Resistance
testing is here. It's commercially available and being used. HIV treatment
guidelines from both the Federal government Health and Human Services
Administration Public Health Service and the International AIDS Society (IAS)
recommend that the use of resistance testing can be useful as a tool in making
certain treatment decisions in individuals with treatment experience who need to
change therapy. But it is important to be aware of the shortcomings of the
tests. At the Resistance Workshop in Spain, I think it was generally agreed that
the use of resistance testing can help a doctor make a better decision in
selecting a regimen for an individual who needs to change therapy because of
virologic failure. But it was also generally agreed that resistance testing is
one of a number of tools that should be used to help select a regimen. It was
also agreed by the test manufacturers and researchers that improvements need to
and will be made to testing and interpretation, as well as application of
results.
A
patient's treatment history and the doctor's judgement of that history are key
elements in making a treatment change. Oftentimes, a treating physicianís good
judgement can determine which drugs a patient may or may not be responsive or
sensitive to. But several studies have shown that using a resistance test
(genotypic or phenotypic) can be helpful in selecting the most effective
regimen.
By
identifying which drugs a person is most sensitive to, resistance testingóand,
perhaps more importantly, proper interpretation of the resultsóshould help
select a regimen with fewer drugs. Using a regimen with fewer drugs promotes
easier tolerability and adherence. So rather than taking a shotgun approach with
six or more drugs, an equally effective or possibly more effective regimen of
four drugs may be selected. Discussion of limitations of resistance testing
received much attention at the Workshop. If
a person has extensive treatment experience, testing may not be as helpful to
selecting a new regimen. For one, many of the drugs the person used have not
been used for a long time, and resistance may not be detectable. Therefore, it
is generally agreed that resistance testing should be performed while the person
is on therapy. But even then, resistance to previously used drugs may not be
apparent. There is a second concern regarding people with extensive drug
experience. People with extensive drug
experience may not be adequately sensitive to any of the available drugs.
Testing may not help these individuals identify an effective regimen.
Several
studies have shown that individuals who were fully sensitive to more drugs did
better. For example, one study showed that individuals fully sensitive to 0 or 1
drugs using resistance testing achieved a transient 1 log viral load reduction
which was gone after 8 weeks on the new regimen. But, individuals who were fully
sensitive to 2 or 3 drugs had a 1.5 log reduction after 8 weeks on the new
regimen, and after 20 weeks on therapy individuals had a 2.5 log reduction.
A
third concern is that resistance testing
may not detect resistance present at low levels in a person's circulating
virus population. Reported at the Resistance Workshop, Allan Hance used
ultra-sensitive detection of minority populations of resistant virus using a
real-time PCR-based method that relies on selective amplification and
quantification of viruses carrying the V82A and/or the L90M mutations in the
protease. Resistant virus was found at low levels. Resistant viral quasispecies
representing <0.1% (mutation L90M) and 1% (mutation V82A) of the total virus
population could be detected. Hance followed four patients undergoing STIs in
whom complete reappearance of wild-type virus was observed by two to three
months. Using conventional genotyping, resistant virus could still be detected
for as long as five months after interruption of therapy, and was undetectable
at three months in only two out of four patients using the ultra-sensitive PCR
method. In the patients with L90M and/or V82A at baseline, resistant virus was
still present at three months in 9 out of 12 cases.
An
additional concern is the ability to
interpret phenotypic resistance fold changes when they are near the
"cut-off". For example, the Virco cut-off between no resistance
and intermediate resistance is a 4.0 fold change in susceptibility while for the
Virologic test the fold change is 2.5. So, how do you judge a 3.0 or 1.8 change
in d4T susceptibility? In addition, we
don't know how to judge resistance to d4T. Below is a study reported on at
the Workshop comparing Virco and Virologic tests and it discusses discordant
results that are close to the cut-offs. Using the current cut-offs there is no
way to judge phenotypic resistance to double PI combinations such as
indinavir-ritonavir, ritonavir-saquinavir or to the new PI ABT-378. Ten or
possibly even twenty-fold decreased susceptibility to ABT-378, may not mean
resistance. But Abbott, the manufacturer of ABT-378, and the two test makers are
in discussions about setting appropriate cut-offs.
It
has been difficult to demonstrate specific genotypic and phenotypic resistance
to d4T and ddI in the clinic. Vincent Calvez reported study findings at the
Resistance Workshop, that modest increases of d4T IC50 and IC90 may change the
viral load response suggesting the need for a new definition of decrease in d4T
susceptibility. For d4T index threshold values higher than 1.8 (threshold of 4.0
is used in Virco test and 2.5 is used in Virologic phenotypic test) the viral
response was very modest (decrease lower than 0.3 log copies/ml).
Another
concern is that some resistance mutations may not have been identified yet.
There may be mutations that can reflect resistance to a drug but if researchers
don't know about it, resistance may not be identifiable. Additionally, two
mutations may have a special relationship that has not yet been identified. For
example, 3TC resistance (M184V) may partially reverse resistance to AZT. Some PI
mutations may have a similar relationship that has not yet been found. 3TC
resistance may increase sensitivity to PMPA, but requires continuing pressure to
select M184.
In
discussing changing therapy for treatment experienced individuals, the recently
updated IAS Guidelines which were published in the Journal for the American
Medical Association said:
Resistance
testing may assist in selecting which drugs should be changed and which could
remainÖ.
Resistance emergence is highly predictive of loss of antiretroviral activity.
In
discussing initial therapy for treatment naÔve individuals, the IAS Guidelines
said, "Choice of a regimen should be individualized based on the strength
of supporting data and on regimen potency, tolerability, adverse effect profile,
likely drug-drug interactions convenience and adherence likel.yhood, potential
for alternative treatment options if the initial regimen fails, and possibly,
baseline resistance testing results."
The
IAS Guidelines also discussed some of the limitations of resistance testing:
costs; some labs may lack quality assurance [a lab should have a track in
delivering reproducibly reliable test results]; the difficulty in interpreting
the test results in its use for a particular person; if a person has used a drug
in the past resistance to that drug may not be detectable with commercially
available testing; resistance in minority species [10-20% of a person's
circulating virus population] is not likely to be detectable. Absence
of detectable genotypic or phenotypic resistance may not provide assurance that
a drug will be active.
Interpretation
of test results requires an appreciation of the range of mutations that are
selected for by various antiretroviral drugs, as well as the potential for
cross-resistance to other drugs conferred by some of these mutations.
Consultation with an expert in HIV drug resistance is encouraged to facilitate
interpretation of genotypic test results.
Transmission
of resistant virus has been reported and "documented," and may be
associated with a suboptimal response to therapy. When deciding to treat during
acute infection, optimization of the initial antiretroviral regimen through the
use of resistance testing is a reasonable although untested strategy. The
Guidelines recommend the use of genotypic testing because the turnaround time in
receiving results should be quicker, but therapy should not be withheld while
awaiting the results of resistance testing.
For
resistance testing, one should use a lab with high expertise in performing
testing (using the kit) and in interpreting the results. It ought to be a lab
that has demonstrated a track record of providing reliably accurate results
reflected in patient treatment decision-making. Immediately following are
studies from Sitges comparing the major resistance tests. The studies point out
that a single resistance test or result report can miss detecting a mutation.
Although any lab test can also make a mistake, and concordance between the two
genotypic testing reported below may appear satisfactory, if you are the patient
and a wrong treatment decision is made based upon inaccurate genotype, that is
unsatisfactory. One pitfall with resistance testing is that the results of
research in populations show benefit, but for an individual, application is more
difficult.
Differences
in Protease and Reverse Transcriptase Sequences between the TruGene HIV-1
Genotyping Kit (Visible Genetics) and the ViroSeq Genotyping System (PE Applied
Biosystems - ABI-).
Assays were performed according to the TruGene HIV-1 genotyping kit and the
ViroSeq genotyping system (Perkin Elmer) package instructions. They only
considered major differences, in which a wild-type genotype was detected by one
method, and a mixed or mutated genotype by the other method.
The
mean total number of mutations associated with resistance to protease inhibitors
and RT inhibitors per sample was 4.52 (±2.04) and 4.70 (±1.95), respectively.
The mean number of primary mutations was 1.85 (±0.92) for PI and 2.54 (±1.00)
for RTI. Among the 96 specimens analyzed, a total of 428 primary and 487
secondary mutations were found with the two methods. Major differences in
primary and secondary mutations have been identified at 14 and 19 amino acid
positions, respectively. Among the 14 discordant (different) primary mutations,
nine were detected by VGI and five by PE. For the protease gene, discordances
concerned only positions 46 (n=2) and 84 (n=1), and for the RT gene, positions
70 (n=1), 74 (n=1), 103 (n=2), 108 (n=1), 184 (n=2), 188 (n=1), 190 (n=2), and
215 (n=1). These 14 major discordances involved 13/96 (13.5%) patients, and in 6
of them genotypic interpretations would have been modified. Among the 19
discordant secondary mutations, five were detected by VGI and 14 by PE. For the
protease gene, discordances concerned positions 10 (n=2), 20 (n=4), 24 (n=3), 33
(n=1), and 54 (n=2), and for the RT gene, positions 41 (n=2), 67 N=2) and 210
(n=3). These 19 discordances involved 16/96 (16.6%) patients, and in two of them
genotype interpretation would have been modified. The authors concluded that the
study results did not favor one kit over the other.
A
Blinded Comparison of Two Sequencing Assays for Resistance Mutations in HIV-1
Protease and Reverse Transcriptase Genes in the GART Study
(CPCRA
046). Marie Hoover
and a U.S. research group including Tom Merigan (Stanford), Doug Mayers (Henry
Ford in Detroit), John Baxter (Robert Wood Johnson Medical School in NJ), and
Mark Winters (Stanford) reported on a study comparing the same two tests. Their
purpose was to determine whether there is agreement in identifying major
protease and RT resistance when utilizing HIV-1 genotyping employing two DNA
sequencing methodologies: (1) an investigator-developed methodology run on ABI
sequencers; (2) Visible Genetics integrated TruGene DNA sequencing system.
The
study examined the previously well-characterized baseline plasma samples
collected from 153 patients. They had all been exposed to at least one protease
inhibitor and at least two NRTIs, and they all participated in the GART study (a
prospective study of the utility of genotypic antiretroviral resistance testing
in patients failing therapy). The specimens were provided to a single lab for
blinded re-analysis using the VGI integrated TruGene DNA sequencing system. The
amino acid sequence differences (as compared to a standard sequence) were
compared with the original genotyping results. Discrepancies in amino acid
mutations were investigated by comparing the DNA sequence chromatographs
generated by the two methods.
Hoover
reported that the comparison showed general agreement between the methodologies
for the major protease resistance mutations (30N 99%, 46I 91%, 46L 92.2%, 48V
100%, 82A 93.5%, 82F 100%, 82T 98.7%, 84V 99.3%, and 90M 91.5%). Similarly,
Hoover reported there was agreement for the major nucleoside RT resistance
mutations (65R 99.3%, 69D 100%, 74V 99.3%, 151M 99.3%, 184V 97.4%, 215F 96.1%,
and 215Y 90.8%). Preliminary analysis of the DNA sequence chromatographs of over
50% of the genotypes with discrepancies revealed that only 17.9% (5/28) of the
discrepancies are due to clear differences in the DNA sequence generated by the
two methods. The remainder of the discrepancies are attributed to reporting
errors (39.3%), and differences in VGI versus GART rules for calling mixtures of
nucleotides (10.7%). A VGI mixture is described as occurring when software
indicates a mixed base on one DNA strand. A GART mixture is described as
occurring when software indicates a mixed base on both DNA strands, and there is
a detection of second base in mixtures of nucleotides (32.1%).
The
authors concluded that overall, the two methods provide similar drug resistance
genotypes.
A
Comparative Analysis of Virco Antivirogram and Virologic PhenoSense Phenotypic
Assays for Drug Susceptibility of HIV-1.
These two phenotypic resistance tests are commercially available either through
LabCorp, the companies themselves, or possibly through other labs. The drug
susceptibilities of the recombinant viruses are measured by different
strategies. Both tests report IC50 values for each drug and fold differences of
these values relative to a reference wild-type (WT) virus. Fold differences
>2.5 and >4.0 are considered evidence of decreased susceptibility for
PhenoSense and Antivirogram, respectively. Shoukat Qari from the CDC reported on
this study, comparing both tests by analyzing 50 coded plasma samples: twenty
from drug-naÔve newly-infected persons, and 30 samples from source patients of
occupational exposures to HIV-1, of which 16 were from drug-experienced
patients. Results of susceptibility to 12-15 drugs were available from both
tests on 38 samples. These results included 246 determinations for nucleoside RT
inhibitors (NRTIs), 111 for NNRTIs, and 172 for protease inhibitors (PI). There
were 488 (92.2%) concordant results, comprising 459 (94%) sensitive results and
29 (6%) with decreased susceptibility. Of
the 41 (7.8%) discordant results reported to have decreased susceptibility by
either test (26 Virco, 15 Virologic), no known primary resistance related
mutations were reported in 40.
The
discordant results comprised 2.6% for NRTI, 3% for NNRTI, and 2.1% for PI. Of
the 41 results, 26 (63.4%) (10 by Virco and 15 by Virologic) were within
one-fold of the assay cut-offs (4.1 - 5.3 fold for Virco, and 2.52 - 3.7 fold
for Virologic). The remaining 15 discordant results were present in 9 samples
and included 5 results by Virco (5.9 - 9.5 fold) and 10 results by Virologic
(4.6 - 18.5 fold).
Dr
Qari pointed out that the significance of some of these discordant results was
unclear, as clinically relevant threshold fold-changes have not been defined.
The authors concluded that despite the use of different testing strategies in
both tests, the data in this study indicate a good correlation between results
from the two tests. At the Workshop it was discussed and generally agreed that
low level NNRTI resistance (<10 fold) may not be significant, particularly if
a person were NNRTI naÔve.
Virtual
Phenotype.
Brendan Larder, with Virco, discussed Virco's genotype interpretation system
called Virtual Phenotype. The Belgium-based Virco has a database of over 30,000
genotypes and 40,000 phenotypes, with 13,000 matched samples. The system uses a
computerized approach to match a genotype sample from a patient with the
genotypes in the system. Upon retrieving phenotypes from the matching samples in
the database, they calculate the phenotype for the submitted patient sample. The
result is a quantified prediction of phenotype sensitivity of the patient's
sample. Larder reported data showing a correlation between Virtual Phenotype and
actual phenotype. He also suggested Virtual Phenotype was a more accurate
predictor of resistance than "rules based" genotype systems that they
compared.
Resistance
Prevalence.
Quest Labs and Stanford University reported from a database (11,000 samples)
which they collected that over a 24-month period, there has been a significant
increase in multi-drug resistance in patient samples submitted for testing to a
clinical lab. Significant increases from 1998 to 1999 were observed in the
percent of samples resistant for the NNRTIs nevirapine (S1 '98: 22.2%; Q4 '99
43%), delavirdine (S1 '98: 19.8%; Q4 '99 39%), and efavirenz (S1 '98: 18%; Q4
'99: 38%). The frequency of the K103N mutation associated with NNRTI resistance
has more than doubled, accounting for NNRTI resistance. Lesser increases were
seen in NRTI and PI resistances. The authors concluded that the development of
new drugs designed specifically to target the most common resistant variants of
HIV-1 RT and protease may prove to be a more effective strategy for combating
the spread of resistant virus.
Resistance
In Newly Infected.
Virologic and Aaron Diamond AIDS Research Center reported on their monitoring of
the incidence of drug resistance among newly infected individuals. They assessed
baseline resistance in all 41 newly infecteds referred to the ADARC during April
1 1999 to March 31 2000. Three patients were identified with significant
phenotypic resistance to any drug, including two cases of multi-drug resistance.
MDR virus number 1 revealed a highly reduced susceptibility to all protease
inhibitors beside amprenavir, as well as 49-fold reduced susceptibility to AZT
(genotype- RT M41L, D67N/S, T69D, V118I, T215Y; protease- K20I, M36I, M46I,
L90M). Number 2 virus: >50 fold reduced susceptibility to all 3 NNRTIs and an
intermediate 6-fold resistance to NFV (genotype: RT K103N, Y181C; protease:
K20I, M36I, L90M). Number 3 virus: high resistance to NVP (10.9-fold) and DLV
(33.6 fold) was detected.
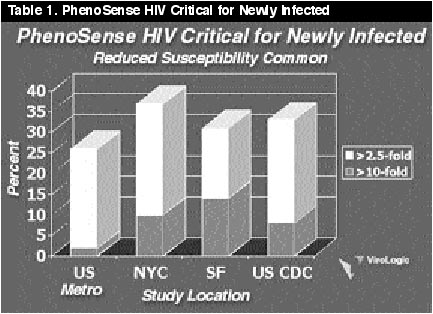
The overall prevalence of any resistance conferring codon changes in RT was 27% (10/36). Substitutions frequently involved in protease gene polymorphisms were L63P/Q/C/S (53%), V77I (28%), L10F/I (13%), I93L (25%), and M36I (15%). L90M was the only primary mutation detected in protease (5%). Hypersensitivity to any NNRTI (<0.5 fold) was seen in six individuals, of whom only two showed classical AZT mutations. The authors concluded that in comparison to previous years they documented increasing prevalence of genotypic altered viruses. Clinical relevance has yet to be established. (See Tables 1 & 2; Much thanks to Virologics, Inc for the use of these slides)

Table
1 shows
the incidence of resistant virus transmitted to newly infected individuals by
certain geographic areas. It appears the incidence varies geographcally.
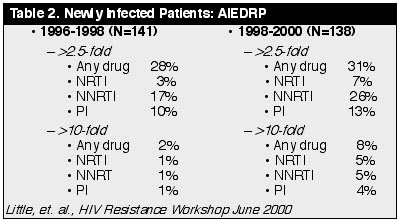
Table
2
indicates that the incidence of transmitted resistant virus has increased in the
last 2 years, suggesting it may continue to increase.