 |
 |
 |
| |
SELECTION AND CHARACTERIZATION OF HEPATITIS C VIRUS REPLICONS RESISTANT TO A POTENT POLYMERASE INHIBITOR A-837093
|
| |
| |
Reported by Jules Levin
AASLD, Oct 27-31, 2006, Boston, MA
L. Lu, P. Krishnan, R. Pithawalla, T. Dekhtyar, T. Ng, W. He, D. Larson, T. Bosse, R. Wagner, D. Kempf, H. Mo, T. Pilot-Matias, A. Molla
Antiviral Research, Global Pharmaceutical Research and Development, Abbott Laboratories, Abbott Park, IL 60064, USA
AUTHOR CONCLUSION
This study suggests that monotherapy with the polymerase inhibitor A-837093
may lead to the development of resistance. However, the lack of crossresistance
may allow this agent to be used in combination with different classes of polymerase inhibitors, protease inhibitors or interferon.
INTRODUCTION
Because of the poor tolerability and the limited efficacy of IFN-based regimens, much effort has been focused on the development of inhibitors of virally encoded enzymes essential for HCV replication. The HCV NS5B RNA-dependent RNA polymerase is a logical target for rational drug screening and design. HCV has a high replication rate in vivo and its polymerase has poor fidelity, thus, it is highly likely that drug-resistant HCV variants will emerge in patients treated with HCV polymerase inhibitors. In fact, in vitro selection of HCV replicons resistant to a number of polymerase inhibitors has been reported. It was found that HCV replicon resistance to polymerase inhibitors has been linked to specific amino acid substitutions within the NS5B polymerase. We and others have found that specific mutation(s) (H95R, N411S, M414L, M414T, Y448H, C451R, and/or G558R) in the NS5B gene led to decreased susceptibility to benzothiadiazines.
Previously, we have reported the antiviral activity and resistance profile of A-782759 (Mo et al., 2005), an analog of the benzothiadiazine series. Further optimization of the A-782759 series has led us to identify compounds A-837093 with significantly improved metabolism and pharmacokinetic properties as well as excellent anti-viral potency on HCV replicon. Because of these features, it is of interest to investigate the resistance profile of this potent inhibitor.
Objective
We undertook this study to characterize the resistance profile of A-837093, a potent HCV RNA dependent RNA polymerase (RdRp) inhibitor.
METHODS
- The anti-HCV activity of A-837093 was determined in HCV replicon cells by the reduction of HCV RNA. HCV RNA was measured by a real-time RT-PCR Taqman assay using primers from the 5' nontranslated region.
- HCV replicon colonies resistant to the inhibitor were selected by treating the HCV sub-genomic 1b-N replicon cells with A-837093 at a concentration 10 times above its EC50 (60 nM) in the presence of 400 _M G418 in the culture medium.
- Colonies were isolated and genotypes of the resistant colonies were determined by sequencing the NS5B polymerase gene which was amplified from HCV RNA by RT-PCR.
- Individual mutations were introduced into a luciferase-expressing replicon by site-directed mutagenesis. Inhibitor susceptibilities of the site-directed mutants were determined using a transient transfection assay by measurement of luciferase activity at day 4. Replication capacities were obtained by comparison of luciferase activities between mutant clones and the wild-type replicon.
RESULTS
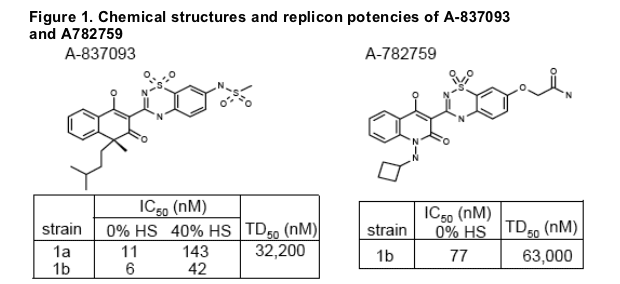
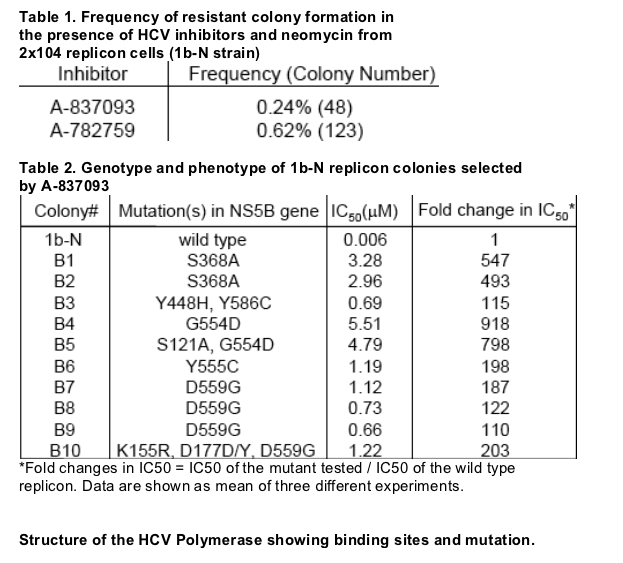
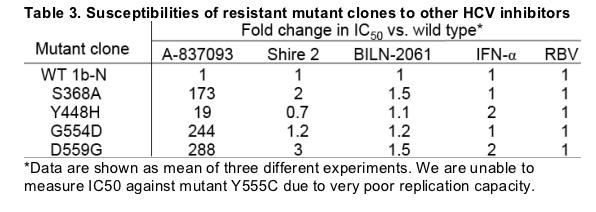
A-837093 displayed excellent activity in HCV replicon with EC50 of 6 nM and 11 nM against HCV genotype 1b-N and 1a-77, respectively (Fig 1). Replicon colonies selected by A-837093 exhibited substantial decreases in susceptibility to A-837093. Genotypic analysis demonstrated that each colony contained one of the following mutations: S368A, Y448H, G554D or D559G in the NS5B gene (Table 2). Molecular clones containing the above single mutations conferred moderate (Y448H, 19-fold) to high-level (200-300 fold) resistance to A-837093. In contrast, all four mutants retained susceptibility to a thiophene-2-carboxylic acid polymerase inhibitor (Shire 2), the HCV protease inhibitor BILN2061 and interferon (Table 3). Mutant replicons showed significantly impaired fitness compared with the wild-type replicon (Fig. 3).
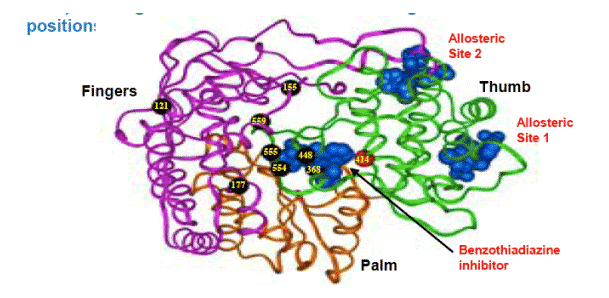
A ribbon representation of the protein in three colors shows the fingers (pink),
palm (brown) and thumb (green) domains. Mutations (from Table 1) are
shown as black balls at the alpha carbons of the specified residues. Residue
414, not identified as a mutation position in this study, is shown as a red ball
for discussion purposes. Residue 586 is located in a C-terminal membrane
anchor domain and is not shown. Space-filling representations of various
inhibitors (blue) are also shown: benzothiadiazine (Tedesco et al., 2006);
allosteric site 1 inhibitor (Wang et al, 2003); allosteric site 2 inhibitor (Di Marco
et al, 2005).
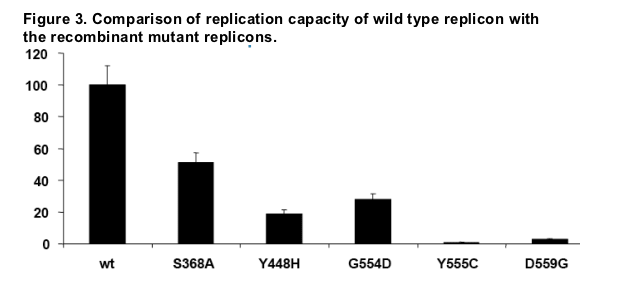
Cells were transfected either with wild-type or with mutant replicon RNA and
luciferase activities were measured at four hours and four days after
transfection (Lu et al., 2004). The replication capacity for each mutant was
calculated by comparing the luciferase activity generated by the mutants to
that generated by wild-type replicon at day 4, after adjusting for minor
differences in transfection efficiencies (the 4 hours luciferase activity). The
data are average of at least two separate experiments with six replicates in
each experiment. The error bars represent the SD.
REFERENCES
Di Marco et al., 2005. Interdomain Communication in Hepatitis C Virus
Polymerase Abolished by Small Molecule Inhibitors Bound to a Novel Allosteric
Site. J. Biol. Chem.. 280:29765-29770.
Lesburg, et al., 1999. Crystal structure of the RNA-dependent RNA polymerase
from hepatitis C virus reveals a fully encircled active site. Nat. Struct. Biol.. 6:937-
43.
Lu, L., et al., 2004. Mutations conferring resistance to a potent hepatitis C virus
serine protease inhibitor in vitro. Antimicrob. Agents Chemother. 48:2260-6.
Mo, H., et al., 2005. Mutations conferring resistance to a hepatitis C virus (HCV)
RNA-dependent RNA polymerase inhibitor alone or in combination with an HCV
serine protease inhibitor in vitro. Antimicrob. Agents Chemother.. 49:4305-14.
Tedesco, R., et al., 2006. 3-(1,1-dioxo-2H-(1,2,4)-benzothiadiazin-3-yl)-4-hydroxy-2(1H)-quinolinones, potent inhibitors of hepatitis C virus RNA-dependent RNA polymerase. J. Med. Chem.. 9;49(3):971-83.
Wang, M., et al., 2003. Non-nucleoside analogue inhibitors bind to an allosteric
site on HCV NS5B polymerase. Crystal structures and mechanism of inhibition. J. Biol. Chem.. 278:9489-95.
|
| |
|
 |
 |
|
|