 |
 |
 |
| |
The composition of dual-mixed HIV-1 populations can be discerned from the level of CXCR4-mediated infectivity
|
| |
| |
Reported by Jules Levin
ICAAC Sept 11-15 2009 San Francisco
Jonathan Toma, Arne Frantzell, Signe Fransen, Jeannette
M. Whitcomb, Christos J. Petropoulos and Wei Huang
Monogram Biosciences
South San Francisco, CA
USA
BACKGROUND
Dual/mixed (DM) viruses represent the majority of CXCR4-utilizing HIV isolates and display a broad range of infectivity in CXCR4+ target cells.
Recent studies have shown that pre-existing minor subpopulations of CXCR4-using viruses can emerge to replace R5 virus populations after initiation of CCR5
antagonist treatment.
To better understand the characteristics and composition of DM populations, we conducted clonal analyses for a panel of DM viruses reflecting the full spectrum of infectivity observed for CXCR4-using viruses.
METHODS
Twenty DM virus populations were selected from routine Trofile patient testing samples. Numerous full-length env clones were isolated from each virus population and evaluated for co-receptor usage.
Env clones were classified as R5-, X4- and dual-tropic using the standard (not Enhanced) Trofile assay; dual clones were further classified as dual-X or dual-R based on the differential production of luciferase, measured as relative light units
(RLU), upon infection of CXCR4+ and CCR5+ target cells. (for the purpose of this study, defined as: dual-R = R5:X4 RLU ratio >10, dual-X = R5:X4 RLU ratio < 10)
env sequences were determined using conventional methods.
RESULTS
The infectivity of 20 DM virus populations in CCR5+ and CXCR4+ target cells varied over a 3-4 log10 range (Figure 1A).
Clonal analyses revealed the presence of both dual and X4 clones in DM viral populations. The proportion of CXCR4-using clones and R5 clones varied widely among different patient samples (Figure 1B).
DM virus populations with higher CXCR4 infectivity contained predominantly X4 and dual-X clones that efficiently use CXCR4. Conversely, virus populations with low CXCR4 infectivity consisted predominately of R5 clones with either minor subpopulations of dual-X clones, or dual-R clones that use CXCR4 inefficiently (Figure 1B).
Dual-X and X4 clones were identified in 19/20 samples analyzed (Figure 1B), including 5/6 of DM viruses with the lowest levels of CXCR4 utilization (pool X4 RLUs <1000).
Tropism predictive algorithms based on substitutions in V3 sequences (PSSM and 11/25KR) accurately assigned less than half of the virus populations as CXCR4-using (Figure 2).
Clones with identical V3 sequences can have varying abilities to infect CXCR4+ cells, which indicates that env sequence changes outside of V3 contribute to CXCR4 use (Figures 3, 4).
Changes in env sequences outside of V3 that alter coreceptor utilization can be subtle (Figure 5).
CONCLUSIONS
This study demonstrates that patient DM virus populations may be comprised of either homogeneous dual-tropic variants or complex mixtures of dual, X4 and R5 variants.
The ability of DM virus populations to infect CXCR4+ cells reflects the proportion of Env clones within the population that efficiently use CXCR4.
R5 and dual clones can share identical V3 sequences, demonstrating that env sequences outside of V3 can impact coreceptor usage.
Our observations further illustrate the diverse composition of DM virus populations as well as the large variation in CXCR4 efficiency of individual clones.
Additional studies are needed to investigate whether or not subpopulations of dual-tropic viruses (e.g. dual-R) within DM virus populations are susceptible to inhibition by CCR5 antagonists, while others (e.g. dual-X) are not.
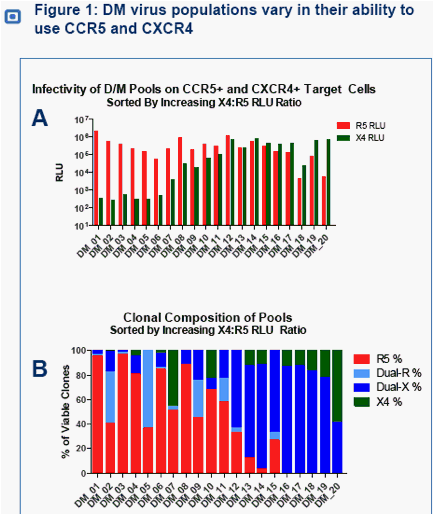

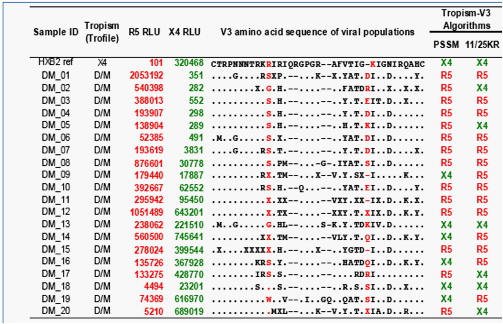



Figure 5: Substitutions in env sequences outside of V3 can confer different levels of CXCR4 use in variants that
share identical V3 sequences
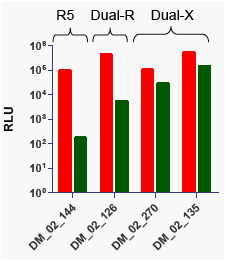

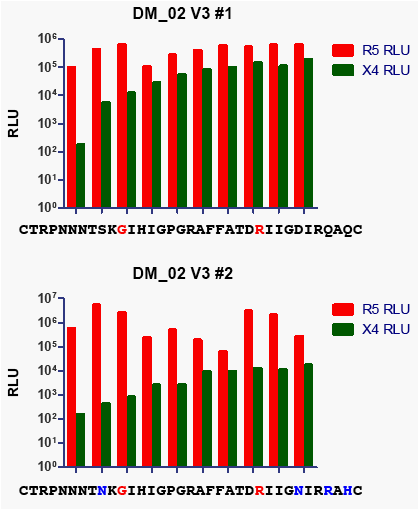
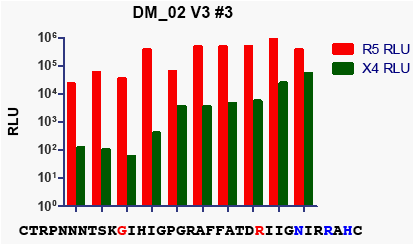
Figure 4: CXCR4 and CCR5 mediated infectivity can
vary among clones with identical V3 sequences (DM-
02 clones)
|
| |
|
 |
 |
|
|