 |
 |
 |
| |
Prediction of HIV Disease Progression Using a Modification of the Enhanced Sensitivity (ES) Trofile Assay
|
| |
| |
Reported by Jules Levin
ICAAC Sept 11-15 2009
San Francisco
Matthew B. Goetz1 , Robert Leduc2, Ann Labriola3, Yolanda Lie4, Jodi Weidler4, Eoin Coakley4, Laura A. Napolitano4, Jay R. Kostman5, Roberta Luskin-Hawk6, for the Long Term Monitoring (LTM) Study, Terry Beirn Community Programs For Clinical Research on AIDS (CPCRA)
1VA Greater Los Angeles Healthcare System, 2University of Minnesota, Minneapolis, MN, USA, 3VA Medical Center, Washington, DC, USA, 4Monogram Biosciences, South San Francisco, CA, USA, 5UPHS/Presbyterian Medical Center, Philadelphia, PA, USA, 6Saint Joseph Hospital, Chicago, IL, USA
AUTHOR DISCUSSION
Identifying samples with the CXCR4 use that was near the limit of detection (NLOD) was relevant to predicting disease progression in this cohort with
chronic untreated HIV-1 infection. Excluding NLOD samples from the DM group defined a group that had a faster rate of HIV disease progression compared to
other individuals. These results were very similar to previous findings in studies utilizing the original Trofile assay. By contrast, no association between CRT and disease progression was found when the DM group included those samples with CXCR4 use NLOD.
Of 278 samples with R5 CRT by the original Trofile assay 24 (8.6%) were
reclassified as DM; this reclassification rate is comparable to that in the MERIT
cohort of treatment-naïve individuals in which 14.7% of R5 tropism samples
were reclassified as DM (6). Of the 24 samples reclassified as DM by ES Trofile,
6 had CXCR4-use that exceeded the low level CXCR4 use control and 18 had
CXCR4-use NLOD.
This is the first study to suggest a possible threshold of CXCR4-use for the Trofile assay with regard to disease progression and contrasts with the impact of the enhanced Trofile reclassification on the response to CCR5 inhibitors (6,
7). Further studies are warranted to explore the possible impact of this threshold effect in other cohorts and to determine whether this threshold effect is associated with quantitative differences in the pool of CXCR4-using virus or qualitative differences in the genetic determinants of CXCR4 utilization (8).
SUMMARY
We have previously demonstrated in the LTM cohort that treatment-naïve individuals with DM tropism experience accelerated disease progression relative to those with R5 tropism as defined by the original Trofile assay.
In the ES Trofile assay this prognostic pattern was confirmed among those with DM tropism where CXCR4-use exceeded that of the low-level CXCR4-use control virus incorporated into the ES Trofile assay. Accelerated disease progression was not seen when CXCR4-use was near the limit of detection (NLOD).
Among 278 samples R5 by the original Trofile, 24 (8.6%) were identified as DM by the ES Trofile assay; Of these 24, 18 had CXCR4 use that was NLOD.
Further studies are necessary to determine if the threshold effect for CXCR4 use and disease progression defined in this cohort is also relevant to other treatment naïve cohorts.
BACKGROUND
Previous studies have shown that untreated chronically HIV-infected persons with CXCR4 coreceptor tropism (CRT) have more rapid disease progression than do persons in whom only CCR5 CRT is detected (1-4). These studies were performed using the original version of the Trofile assay which is validated to detect X4 variants comprising 10% of the viral population
The enhanced sensitivity (ES) Trofile assay is validated to detect X4 variants present at levels >/= 0.3% in vitro (5). The improved sensitivity of the Trofile assay\ has been shown to better identify candidates for CCR5 antagonist therapy (6,7).
Herein we compare the relationship between viral tropism and HIV disease
progression defined by the original vs the ES Trofile assay.
OBJECTIVES & METHODS
· Determine the distribution of CRT using the ES Trofile assay
· Determine the relationship of CRT with the time to < 350 CD4+ cells/µL,
initiation of therapy or death
· Compare the distribution and prognostic value of the detection DM/X4 CRT by the original vs the ES Trofile assays
· Assess the prognostic value of DM/X4 CRT as determined by differing
thresholds of CXCR4 CRT signal strength ES Trofile assays
All samples were from the CPCRA Long Term Monitoring (LTM) Treatment-
Naïve cohort in which outcomes in >12 year-old, antiretroviral- naïve, HIV-infected patients who gave written informed consent were prospectively
monitored. There were no exclusion criteria. Eligibility also required: ≥ 4
months of follow-up during which antiretroviral therapy was not initiated, a
baseline CD4+ cell count ≥450 cells/uL, a baseline viral load >/=1,000 HIV-1 RNA copies/mL and successful prior performance of the original Trofile assay.
The primary analyses explored the impact of identifying samples with CXCR4 use 'near the limit of detection' (NLOD) as defined by a 'low-level CXCR4 use' (LLXC) control incorporated into the ES Trofile assay.
Two groups with DM CRT were defined 1) Any CXCR4 CRT including NLOD
CXCR4 CRT and 2) CXCR4 use but excluding those with NLOD CXCR4 CRT.
RESULTS
359 of the 1090 LTM patients met the entry criteria for analysis of CRT. The
original Trofile assay was successful in 314 specimens; 18 persons did not
have available plasma and the assay was unsuccessful in 27 other samples.
The ES Trofile assay was successful in 310 of these 314 samples.
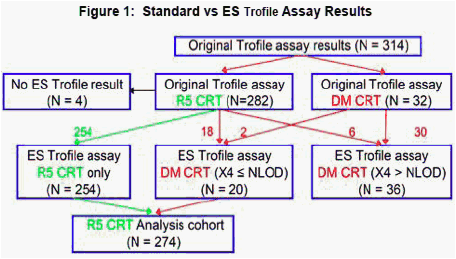
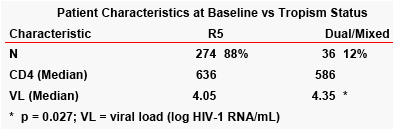
Whereas the original Trofile assay detected dual or mixed (DM) CRT in 32/314 samples, the ES Trofile assay detected DM CRT in 56/310. Excluding the 20 samples with CXCR4-use NLOD defined 36/310 samples with CXCR4 use. Notably all samples with CXCR4 use by the original Trofile assay also
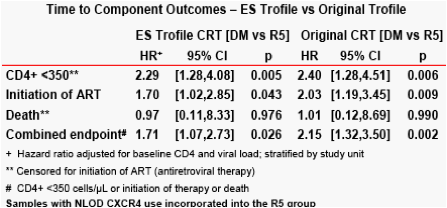
demonstrated CXCR4 use by the enhanced Trofile assay. The sample flow is shown in figure 1. The analyses summarized here incorporate those samples with NLOD CXCR4 use into the R5 group
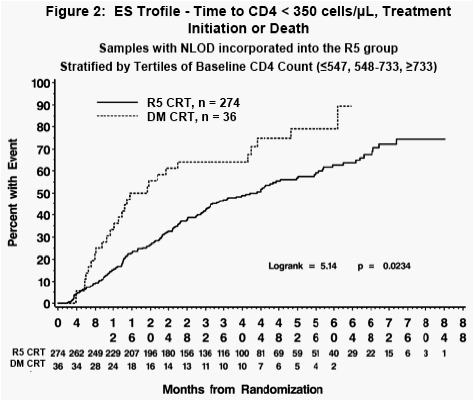
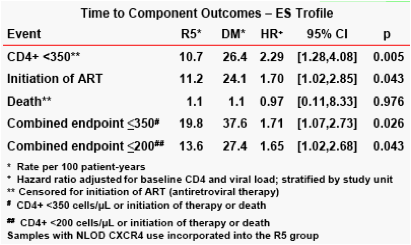
183 patients reached the primary endpoint. The distribution of first events
was as follows: reaching a CD4+ count < 350 cells/µL (n=110); initiation of
antiretroviral therapy (n=64); and death (n=8).


References
1. Waters L, et al. Clin.Infect.Dis. 46, 1617-1623 (2008).
2. Daar ES, et al. Clin.Infect.Dis. 45, 643-649 (2007).
3. Shepherd J, et al. J Infect.Dis. 198, 1104-1112 (2008).
4. Goetz MB, et al. J.Acquir.Immune.Defic.Syndr. 50, 259-266 (2009).
5. Reeves JD, et al. J Viral Entry 3, 94-102 (2009).
6. Saag M, et al. 48th Annual ICAAC/46h Annual Meeting IDSA. 2008; H-1232a.
7. Su Z, et al. 48th Annual ICAAC/46th Annual IDSA Meeting. 2008; H-895
8. Huang W et al. J Virol. 82, 5584-5593 (2008).
|
| |
|
 |
 |
|
|