 |
 |
 |
| |
A high barrier to resistance may contribute to the robust antiviral effect demonstrated by R1626 in HCV genotype 1-infected treatment-naïve patients
|
| |
| |
Reported by Jules Levin
AASLD, Nov 2-6, 2007, Boston, MA
Sophie Le Pogam, Amritha Seshaadri, Alan Kosaka, Steven Hu, Hyunsoon Kang, Julian Symons, Klaus Klumpp, Nick Cammack and Isabel Nájera
Roche Palo Alto LLC, 3431 Hillview Avenue, Palo Alto, CA 94304, USA
Author Conclusions
R1626 exhibited a strong HCV antiviral effect in monotherapy and combination therapy.
The absence of evidence for phenotypic and genotypic changes in patients with viral rebound demonstrates a lack of selection of viral resistance to R1626 after up to 4 weeks of treatment
The viral load rebound observed in a few patients treated with R1626 was likely due to inadequate drug levels (all were from the lowest dose arm of the monotherapy study or following drug dose reduction or discontinuation in patients in the combination study)
Despite the inadequate plasma levels of R1479 in the studied patients (that could represent an opportunity for the selection of drug resistant variants), there was no selection of viral resistance to R1626, neither at a population level nor at low level in the quasispecies, as evaluated through the extensive study of molecular clones
Sequence analysis revealed a lack of pre-existing amino acid substitutions at baseline that could be a predictor of treatment outcome (response or failure)
Given that in vitro studies have shown the poor replication capacity of S96T and S96T/N142T R1479-resistant replicons,2 the lack of selection of viral resistance after up to 4 weeks of treatment may indicate that R1626 exhibits a high genetic barrier to selection of viral resistance
R1626, appears to be a promising agent for treatment of patients chronically infected with HCV genotype 1
Introduction
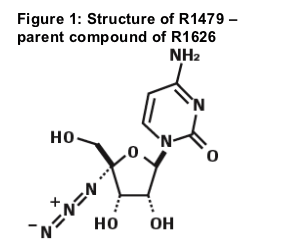
R1479 (Figure 1), Parent compound of R1626 is a potent and highly selective inhibitor of NS5Bdirected hepatitis C virus (HCV) replication in vitro1
In vitro studies have identified NS5B polymerase amino acid substitutions, S96T or S96T/N142T, that result in 4-5-fold reduced sensitivity to R1479 as determined by the HCV replicon assay2
These mutations resulted in a ∴95% reduction in replication capacity compared to the wild type replicon2
R1626 has been shown to be a potent inhibitor of HCV RNA replication in two clinical studies:
1) 14-day monotherapy and 2) 28-day combination therapy with peginterferon alfa-2a [40KD] (PEGASYS) and ribavirin (COPEGUS)3,4
Viral load rebound during treatment was observed in 3 patients from the monotherapy study and in 8 from the combination therapy study
Three patients from the lowest dose of the monotherapy study did not respond to R1626 treatment
We evaluated the potential for in vivo resistance selection in these two clinical studies
Objective
To investigate whether the viral load rebound observed in some of the patients from the R1626 monotherapy or combination therapy studies was due to the emergence of resistance during treatment
To investigate whether the lack of response to R1626 in the monotherapy study observed in 3 patients was due to the pre-existence of resistance mutations in their NS5B coding region
Identify any possible predictors in the NS5B coding region of treatment response or failure
Methods
Patients
Serum samples from patients involved in two multicenter, randomized clinical studies designed to evaluate the safety, pharmacokinetics and antiviral activity of R1626 in treatment-naïve patients chronically infected with HCV genotype (GT) 1 were used to evaluate the in vivo resistance selection:
-- Study 1 was a placebo-controlled, multiple ascending dose, Phase 1b study with 47 patients who received oral R1626 at doses of 500 mg, 1500 mg, 3000 mg or 4500 mg or placebo twice daily (bid) for 14 days with 14 days of follow-up
-- Study 2 was a Phase 2a study that investigated R1626 in combination with PEGASYS and COPEGUS. 103 patients were randomized to: R1626 (1500 mg bid) plus PEGASYS (Dual Low); R1626 (3000 mg bid) plus PEGASYS (Dual High); R1626 (1500 mg bid) plus PEGASYS and COPEGUS (Triple Low); or PEGASYS plus COPEGUS (standard of care) for 4 weeks
Viral resistance evaluation
The viral load profiles of all participants were assessed for a viral rebound phenotype defined as:
-- A ≥ 0.5 log10 IU/mL increase of viral load from nadir during treatment with R1626, where nadir is defined as a ≥ 0.5 log10 IU/mL decrease from baseline (lowest point)
Selection of patient samples for resistance monitoring
-- Samples from patients who experienced a viral load reduction ≥ 0.5 log10 IU/mL followed by a viral load rebound were taken before treatment (baseline) and at the time point after the viral load rebound was observed, which corresponded to the end of treatment point for most patients. For 2 patients, the rebound occurred earlier and the end of treatment time point was also included in
the study
--Measurement of sensitivity to R1479 in the HCV NS5B phenotypic assay
HCV RNA was extracted from patients' serum and the NS5B coding region was amplified and cloned into replicon vectors (GT 1a samples in GT 1a H77 replicon and GT 1b samples in GT 1b Con1 replicon). Ninety-six clones of each sample were pooled to represent the viral genetic diversity present in the patients
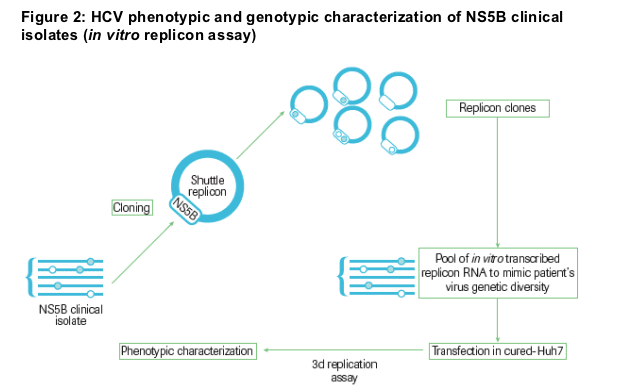
In vitro transcribed replicon RNA containing the NS5B region from the clinical isolates was transfected into cured-Huh7 cells
-- R1479 (or medium as a control) was added 24 hours post-transfection in 3-fold dilutions at a final DMSO concentration of 1%
-- Firefly luciferase reporter signal was read 72 hours after addition of compounds and the IC50 values were assessed as the compound concentration at which a 50% reduction in the levels of firefly luciferase reporter was observed as compared to control samples in the absence of compound
Sequencing of NS5B coding region
-- The entire NS5B polymerase coding region was sequenced
-- Detailed analysis of the NS5B viral quasispecies was performed by sequencing of the 96 molecular clones included in the pool to represent the viral genetic diversity
Results
R1626 showed a time- and dose-dependent HCV RNA reduction
in monotherapy and combination therapy
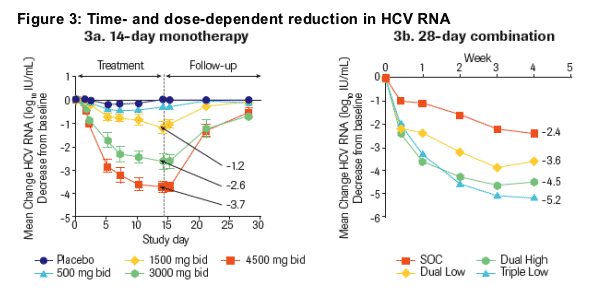
Study 1: Time- and dose-dependent reductions in HCV RNA levels were achieved with mean HCV RNA reductions of up to 3.7 log10 IU/mL following 2 weeks of monotherapy (Figure 3a)
Study 2: Time- and dose-dependent reductions in HCV RNA levels were achieved with mean reductions of 5.2 log10 IU/mL in patients treated with Triple Low, 4.5 log10 IU/mL in patients treated with Dual High, 3.6 log10 IU/mL in patients treated with Dual Low and 2.4 log10 IU/mL in patients treated with SOC after 4 weeks (Figure 3b)
R1626 resistance evaluation
Among all 150 participants from all dose groups in both studies, 11 patients experienced a viral load rebound as follows:
-- 3 patients from the 500 mg bid dose group in the monotherapy study (Patients 1-3 in Table 1)
-- 8 patients from the combination therapy study:
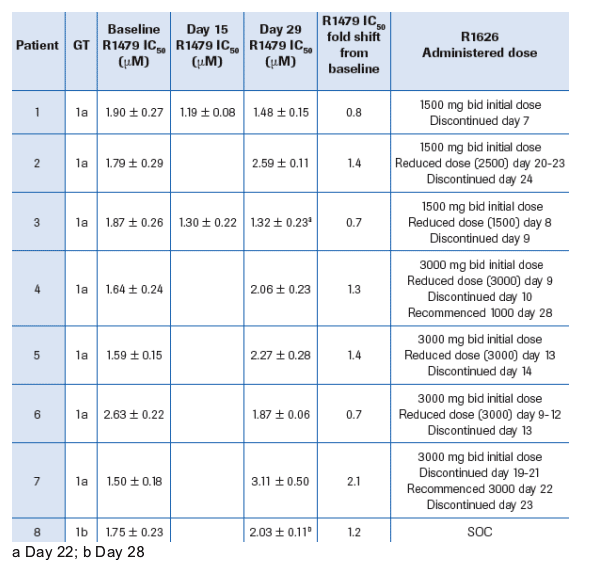
-- 3 in the R1626 1500 mg bid plus PEGASYS group (patients 1-3 in Table 2)
-- 4 in the R1626 3000 mg bid plus PEGASYS group (patients 4-7 in Table 2)
-- 1 in the PEGASYS plus COPEGUS (standard of care) group (patient 8 in Table 2)
Among all participants from all dose groups in both studies, 3 patients did not respond to the treatment in the 500 mg bid dose group of the monotherapy study
No reduction in sensitivity to R1479 after up to 28 day treatment
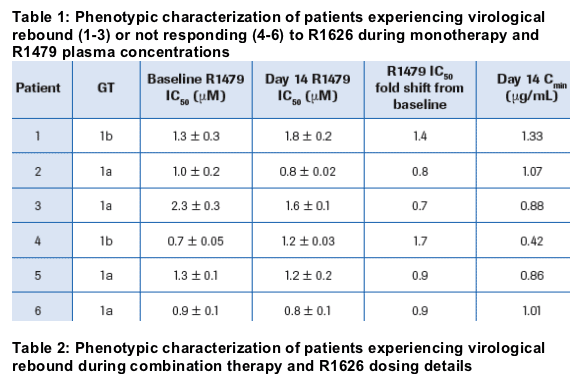
Phenotypic characterization of the clinical isolates from patients receiving R1626 monotherapy that experienced a viral load rebound showed no change in susceptibility to R1479, with a 0.7-1.4-fold shift in IC50 at Day 14 compared to baseline (patients 1, 2 and 3 in Table 1); these values are well within the 2.6-fold variability of the assay compared to baseline sample and compared to reference controls (HCV GT 1a H77 and GT 1b Con1 reference strains, data not shown)2
Phenotypic characterization of the clinical isolates from patients receiving R1626 combination therapy study showed no change in susceptibility at any of the time points compared to their respective baseline or to the reference controls (Table 2)
Phenotypic characterization of the clinical isolates from patients receiving R1626 monotherapy that did not respond to the treatment showed no change in susceptibility to R1479 (patients 4, 5 and 6 in Table 1)
The patients who had viral rebound in the monotherapy study had low levels of R1479 in plasma as shown by the pharmacokinetic data presented in Table 1. Cmin values were lower (0.42 to 1.33 _g/mL) than the overall Cmin range (0.86 to 5.98 _g/mL)
All patients who showed viral rebound on combination therapy had discontinued therapy or had dose reductions (Table 2)
No known R1479 resistance mutations nor other common
substitutions in any of the patients that showed a viral load rebound during treatment or a non response to R1626
In the monotherapy study, NS5B sequence comparisons between baseline and day 14 time points revealed no pre-existing amino acid substitutions (S96T or S96T/N142T) known to be responsible for in vitro resistance to R1479 nor common amino acid changes across the three patients that showed a viral load rebound before the end of treatment
Likewise, NS5B sequence comparison between baseline and treated time points from the combination therapy study revealed no known R1479 resistance mutations nor any other common amino acid substitutions selected upon treatment. Detailed analysis of the quasispecies of all
rebound samples (∴1000 NS5B molecular clones) did not show any known R1479 resistance mutations at baseline or during treatment at low level (sensitivity of 1%)
In the patients that did not respond to R1626 monotherapy, no known R1479 resistance mutations nor other common substitutions were found. Detailed analysis of the NS5B viral quasispecies of the baseline sample showed no pre-existing amino acid substitution responsible for resistance to R1479
No pre-existing amino acid substitutions were observed at baseline that could be a predictor of treatment outcome (response or failure)
References
1. Klumpp K, et al. The novel nucleoside analog R1479 (4'-azidocytidine) is a potent inhibitor of NS5B-dependent RNA synthesis and hepatitis C virus replication in cell culture. J Biol Chem 2006;281:3793-9
2. Le Pogam S et al. In vitro selected Con1 subgenomic replicons resistant to 2a-C-Methyl-Cytidine or to R1479 show lack of cross resistance. Virology 2006;351:349-59
3. Roberts S et al. Results of a Phase 1B, multiple dose study of R1626, a novel nucleoside analog targeting HCV polymerase in chronic HCV genotype 1 patients. Hepatology 2006;44(Suppl 1):692A
4. Pockros PJ et al. Robust synergistic antiviral effect of R1626 in combination with peginterferon alfa-2a (40KD), with or without ribavirin
-- Interim analysis results of Phase 2a study. AASLD 2007 abstract
>
|
| |
|
 |
 |
|
|