 |
 |
 |
| |
BI 201335, a potent HCV NS3 protease inhibitor, in treatment-naïve and -experienced chronic HCV genotype-1 infection: genotypic and phenotypic analysis of the NS3 protease domain
|
| |
| |
Reported by Jules Levin
EASL 2009 April 23-26 Copenhagen
G. Kukolj1, Y. Benhamou2, M.P. Manns3, M. Bourliere4, S. Pol5, M. Schuchmann6, M. Cartier1, D. Huang7, L. Lagace1, G. Steinmann8, J.O. Stern7
1Biological Sciences, Boehringer Ingelheim (Canada) Ltd, R&D, Laval, Canada; 2Service hepato-gastroenterologie, Batiment des cliniques medicales, Hopital Pitie Salpetriere, Paris, France; 3Zentrum Innere Medizin, Abteilung für Gastroenterologie, Hepatologie und Endokrinologie, Medizinische Hochschule Hannover, Hannover, Germany; 4Service d'Hepato-Gastro-Enterologie, Hopital Saint Joseph, Marseille; 5Pole Medico Chirurgical d'Hepato-Gastro-Enterologie, Unite d'Hepatologie, Hopital Cochin, Paris, France;6I. Med. Klinik und Poliklinik, Klinikum der Johannes Gutenberg, Universitat Mainz, Mainz, Germany; 7Virology, Boehringer Ingelheim Pharmaceuticals, Inc., Ridgefield, USA; 8Clinical Research, Virology, Boehringer Ingelheim Pharma GmbH & Co. KG, Biberach, Germany
AUTHOR DISCUSSION AND CONCLUSIONS
HCV NS3 variants that confer resistance to BI 201335 were selected during treatment. These substitutions do not alter the sensitivity to PegIFN/RBV, and the majority of treatment-naïve patients with resistant virus subsequently displayed antiviral responses during combination therapy
The predominant GT 1a resistance mutations encoded a R155K substitution, whereas GT 1b virus mainly encoded changes at D168, with valine as a predominant substitution
The differing persistence of BI 201335 resistance mutants in long-term follow-up among samples in this study suggest D168V variants are less fit in the absence of drug pressure than R155K variants
HCV replicon shuttle vectors have been developed for the phenotyping of the NS3 protease domain from clinical isolates and allow for the identification of variants with altered sensitivity to HCV NS3 protease inhibitors over a large dynamic range
In the treatment-experienced cohorts there was an inverse relationship between the dose of BI 201335 and the number of patients in whom resistant virus emerged. Overall, the results of this study support the further evaluation of BI 201335 in combination with antiviral agents for the treatment of chronic HCV infection
Abstract
Background: BI 201335, a potent and specific HCV NS3/4A protease inhibitor, has been studied with chronic genotype (GT) 1 HCV infection for 14-day monotherapy in treatment-naïve patients followed by combination with PegIFN/RBV for an additional 14 days; and in treatment-experienced patients for 28 days as combination therapy with PegIFN/RBV. All dose groups achieved median viral load reductions of ≥3 log10. On-treatment viral rebound was observed in most patients receiving monotherapy, but in only 3/19 treatment-experienced patients receiving triple combination therapy with lower dosages of BI 201335. Genotypic and phenotypic characterization of viral isolates from this study has been performed.
Methods: HCV RNA was collected and extracted from 53 patients at baseline, days 14 and 28, and during follow-up. Population sequencing was performed on the complete NS3/4A region. Clonal sequencing permitted the quantification and identification of major and minor variants (LLOQ: 5%) in the NS3 protease domain. Chimeric HCV sub-genomic replicons were constructed to determine BI 201335 EC50 values against patient-derived protease sequences.
Results: Population sequencing of baseline isolates revealed one encoding an NS3 V/I170T substitution that conferred an 8-fold reduction in BI 201335 sensitivity (increased EC50) relative to the subtype reference. Mean EC50 values for the other GT 1a (10 ± 8 nM) or 1b (9 ± 4 nM) baseline-derived samples were within 3-fold of the reference values. The predominant GT 1a resistance mutations in on-treatment viral rebound samples encoded an R155K substitution, whereas GT 1b viruses mainly encoded changes at D168, with valine as a predominant substitution. R155K variants conferred reductions in sensitivity to BI 201335 with EC50 values of 1.8-6.5 µM, whereas the EC50s for D168V variants were 3.6-15 µM. Rapid changes in mutant prevalence at the end of dosing suggest that D168V variants are significantly less fit than R155K variants.
Conclusion: HCV NS3 variants that confer resistance to BI 201335 were selected during treatment; these variants do not alter the sensitivity to PegIFN/RBV as the majority of treatment-naïve patients with resistant virus subsequently displayed antiviral responses during combination therapy.
Introduction
· Hepatitis C virus (HCV) serine protease NS3/4A is required for the maturation of the HCV polyprotein during viral replication and is therefore a key molecular target for drug development
· BI 201335 is a specific and reversible peptidomimetic inhibitor of HCV NS3/4A protease. HCV subgenomic replicon testing resulted in EC50 values of 3.1 and 6.5 nM against subtype 1b (Con-1) and 1a (H77), respectively
· As reported previously,1,2 one of the objectives of the first clinical trial of BI 201335 in HCV-infected patients was to study its antiviral activity in treatment-naïve and -experienced, chronic, genotype (GT) 1 HCV-infected patients. Two treatment regimens were investigated:
- 14-day monotherapy (20 mg, 48 mg, 120 mg and 240 mg) in treatment-naïve patients followed by combination with pegylated interferon-a (IFN-a)-2a plus ribavirin (PegIFN/RBV) for an additional 14 days
- 28-day combination therapy (48 mg, 120 mg and 240 mg BI 201335) with PegIFN/RBV in treatment-experienced patients (triple-combination therapy)
· In these studies, BI 201335 was safe, well tolerated and induced a rapid and steep dose-related virologic response within 2 to 4 days of initiation of monotherapy or combination therapy. All groups achieved median maximal viral load (VL) reductions of ≥3 log10. On-treatment viral rebound was observed in most patients receiving monotherapy, but in only 3/19 treatment-experienced patients receiving triple-combination therapy
· Here we report investigational procedures for the genotyping and phenotyping of clinical isolates of HCV, to monitor the emergence of viral resistance during the administration of HCV NS3 protease inhibitors
· In particular we highlight:
-- Interim resistance data
- Viral evolution during BI 201335 treatment and following the end of dosing
METHODS
· Viral RNA extraction and PCR amplification
- Viral RNA was isolated from the plasma of 53 HCV-infected patients at baseline, Days 14 and 28, and during follow-up
- A DNA fragment containing the complete NS3-NS4A region was reverse-transcribed and amplified via PCR
- Two PCR products containing either the entire NS3/NS4A domain or only the NS3 protease domain were generated in a second-round PCR
- The lower limit of detection of the RT-PCR amplification method restricted the analysis to patient samples with a VL ≥1,000 IU/mL
· Sequence analysis
- The NS3/NS4A-containing DNA product was used for direct population-based sequencing of the NS3-NS4A region
- The NS3 protease-containing DNA fragment was used to generate the clonal sequences of the NS3 protease region
-- The resulting sequences were compared to reference sequences according to their respective subtypes; AF009606 served as the reference for subtype 1a and AJ238799 for subtype 1b
· Drug sensitivity assays3
- The first-round PCR products, from HCV-infected patient plasma samples, were used to amplify DNA fragments containing the NS3 protease domain
- NS3 amplicons were ligated into a HCV replicon shuttle vector (pIT2) containing a luciferase reporter gene, and reconstituted plasmid DNA was used to generate HCV subgenomic replicon RNA transcripts
- In vitro transcribed RNA was electroporated into Huh-7.5 cells and luciferase activity was measured 96 hours post-transfection, as a marker for HCV RNA replication
- Susceptibility to BI 201335 was revealed by quantification of luciferase activity across a range of inhibitor concentrations (added 24 hours post-transfection)
-- The concentration of BI 201335 giving 50% inhibition of HCV RNA replication (EC50) was determined
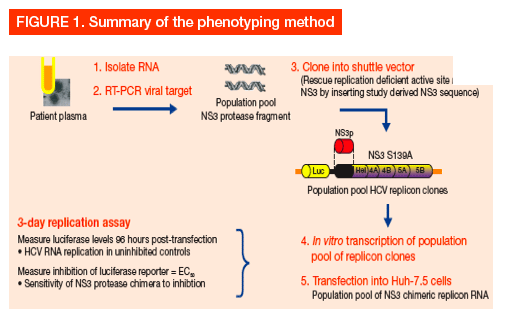
Results
In vitro drug susceptibility of viral NS3 protease
· The chimeric HCV subgenomic replicons constructed from baseline NS3 protease sequences were used to evaluate drug susceptibility with BI 201335 and IFN-a (Figure 2)
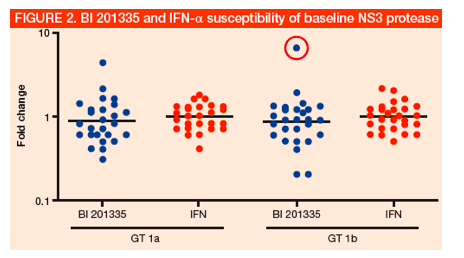
· The average susceptibility of each tested baseline NS3 protease sample to BI 201335, relative to the wild-type (WT) (Con-1b) replicon control values, is depicted as a fold-change. Mean EC50 values for GT 1a (10 ± 8 nM) or 1b (9 ± 4 nM) baseline-derived samples, represented as horizontal lines, were within three-fold of the values obtained with non-chimeric reference replicons
· IFN-a was used as a control for the intrinsic variability of this assay. The mean EC50 value for IFN-a among GT 1a and 1b NS3 protease chimeras was 0.19 ± 0.06 IU/mL and 0.19 ± 0.07 IU/mL, respectively
· The circled sample represents the largest shifting GT 1b NS3 sequence. The genotypic information for this baseline viral sample was compared to the respective reference sequence
- One particular amino acid change in the protease domain was V170T, a locus that was previously demonstrated to reduce the susceptibility of other HCV protease inhibitors4,5
-- Initial BI 201335 antiviral response (first slope decline) in the first-dose group (20 mg QD in treatment-naïve) was lowest for the V170T harboring variant (see 20N3 in Figure 3)
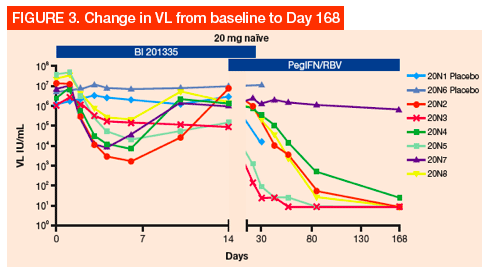
In addition to evaluating all baseline samples, we amplified the NS3 samples isolated from the rebound phase, as well as follow-up. This system was suitable for phenotypic studies of the drug susceptibility for a wide range of GT 1 NS3 sequences, in both the treatment-naïve (Figure 4) and treatment-experienced (Figure 5) dose groups
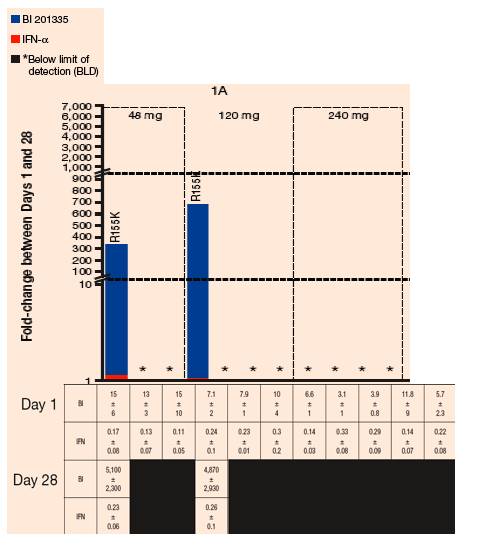
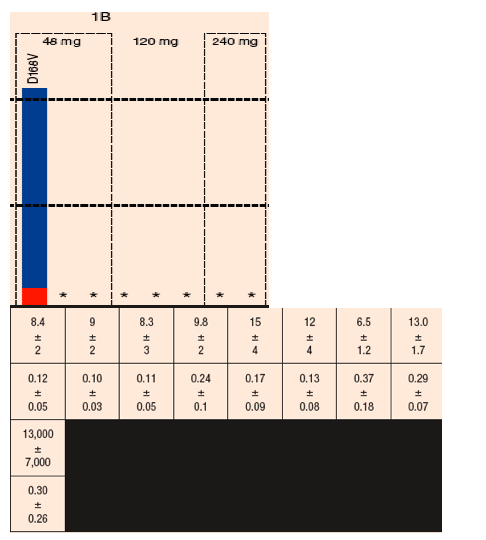
FIGURE 4. Sensitivity of chimeric NS3 replicons to BI 201335 and IFN-a: Fold-change in EC50 between Days 1 and 14 among samples from the treatment-naïve dose groups
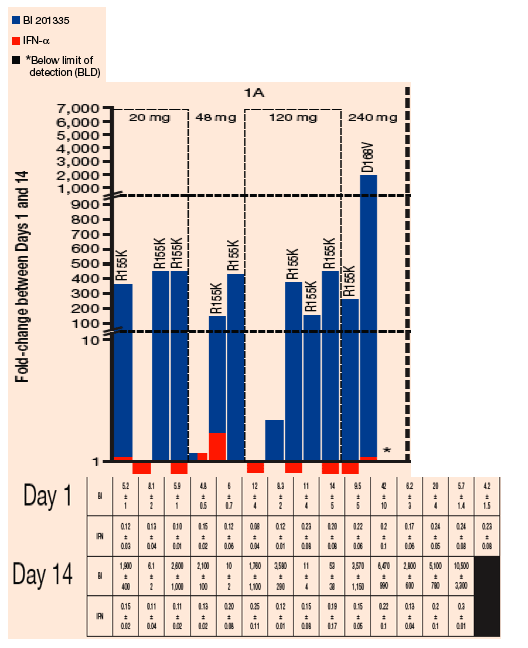
The predominant GT 1a resistance mutations in on-treatment viral rebound samples encoded an R155K substitution - R155K variants conferred reductions in sensitivity to BI 201335 with the range of EC50 values of 1.8-6.5 µM
The GT 1b viruses mainly encoded changes at D168, with valine as a predominant substitution- EC50 values for D168V variants ranged 3.6-15 µM
This profile may be attributable, in part, to a different mutational barrier to resistance at the R155 codon in GT 1a(a single nucleotide transition changes the codon to a lysine) versus GT 1b (that requires two nucleotide substitutions to encode a change to lysine)
One GT 1a sample in the 240 mg dose group encoded for a D168V change at Day 14 (Figure 4); notably, the R155 codon in this variant resembles a 1b subtype
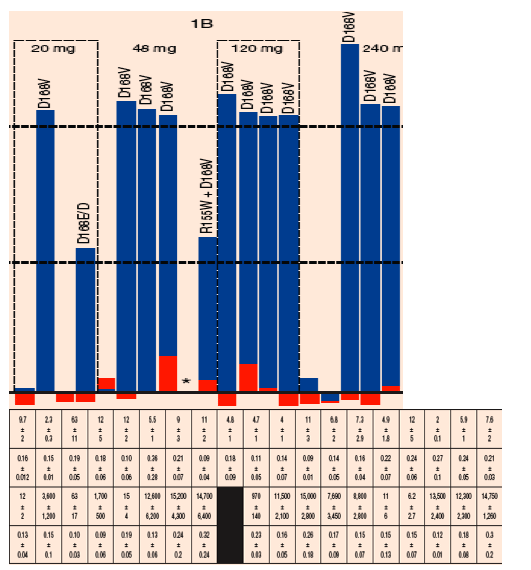
FIGURE 5. Sensitivity of chimeric NS3 replicons to BI 201335 and IFN-a: Fold-change in EC50 between Days 1 and 28 among samples from the treatment-experienced dose groups
Longitudinal clonal sequence analysis
Key resistant mutants were monitored by clonal sequence analysis6 at the indicated timepoints in follow-up toBI 201335 treatment. Figures 6-8 depict three typical examples highlighting changes at NS3 amino acids 155 or168 with a prevalence greater than 2%
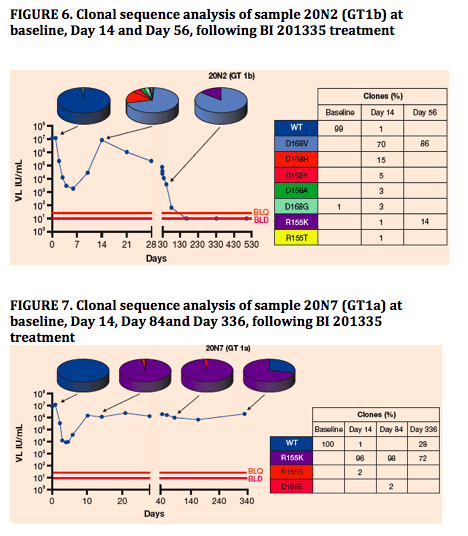
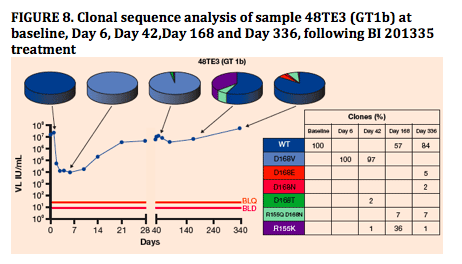
In individual longitudinal analysis, the changing proportions of mutant clones during the follow-up to BI 201335 treatment suggest that D168V variants are substantially less fit in the absence of drug pressure than R155K variants
No D168V variants have been detected in any samples from this study beyond Day 84 due to replacement with WT (or other variants), or to virologic response during SOC treatment
REFERENCES
1. Manns MP, et al. Safety and antiviral activity of BI 201335, a new HCV NS3 protease inhibitor, in treatment-naïve patients with chronic hepatitis C genotype 1 infection given as monotherapy and in combination with peginterferon alfa-2a (P) and ribavirin (R). The 59th Annual Meeting of the American Association for the Study of Liver Diseases (AASLD); San Francisco, CA, USA; 2008. Abstract 1849.
2. Manns MP, et al. Safety and antiviral activity of BI 201335, a new HCV NS3 protease inhibitor, in combination therapy with peginterferon alfa-2a (P) and ribavirin (R) for 28 days in P+R treatment-experienced patients with chronic hepatitis C genotype 1 infection. The 59th Annual Meeting of the American Association for the Study of Liver Diseases (AASLD); San Francisco, CA, USA; 2008. Abstract 1882.
3. Lagace, et al. Development of an HCV sub-genomic replicon based system for phenotyping the NS3 protease from clinical isolates. The3rd International Workshop on Hepatitis C- Resistance and New Compounds; Boston, MA, USA; 2008. Abstract 11.
4. Zeuzem S, et al. Anti-viral activity of SCH 503034, a HCV protease inhibitor, administered as monotherapy in HCV-1 patients refractory to pegylated interferon (PEG-IFN-a). Hepatology. 2005;42(Suppl. 1):233A.
5. Tong X, et al. Identification and analysis of fitness of resistance mutations against the HCV protease inhibitor SCH 503034. Antivir Res. 2006;70:28-38.
6. Sarrazin C, et al. Dynamic hepatitis C virus genotypic and phenotypic changes in patients treated with the protease inhibitor telaprevir. Gastroenterology. 2007;132:1767-1777.
|
| |
|
 |
 |
|
|