 |
 |
 |
| |
Bulk sequence-detectable resistance mutations in peripheral RNA following single-dose nevirapine are associated with poorer treatment responses but do not adequately explain treatment failure
|
| |
| |
Reported by Jules Levin
18th Intl HIV Drug Resistance Workshop
June 9-12 2009 Ft Myers Florida
JA Johnson1, JF Li1, C Yang2, I Zulu3, L Njobvu4,
J Stringer4, V Shanmugam2, A McNulty2,
E Lehotzky2,5, O Bolu2, P Peters1, S Pathak1,
W Heneine1 and P Weidle1
1Division of HIV/AIDS Prevention, CDC, Atlanta, GA, USA
2Global AIDS Program, CDC, Atlanta, GA, USA
3Global AIDS Program, CDC, Lusaka, Zambia
4UAB Center for Infectious Disease Research, Lusaka, Zambia
5Emerging Infectious Diseases Fellow, CDC/APHL, Atlanta, GA, USA


ABSTRACT
BACKGROUND: Sensitive drug resistance testing has revealed that drug resistance emerges in the majority of women exposed to single-dose nevirapine (sdNVP), raising concern for future treatment responses. We examined sdNVP-associated mutations by bulk sequencing and sensitive real-time PCR to determine variant levels and patterns associated with poor treatment response.
METHODS: Viral RNA from 162 sdNVP-exposed and 265 sdNVP-unexposed Zambian women were baselinetested for nevirapine-associated mutations prior to beginning nevirapine/efavirenz-based antiretroviral therapy. Bulk sequencing evaluated reverse transcriptase codons 1-251 and real-time PCR screened only sdNVP-exposed women for K103N, V106M, Y181C and G190A. Presence of baseline majority and/or minority mutations and time since sdNVP exposure were assessed against treatment failure (VL>400 copies/ml) at 24 or 48 weeks.
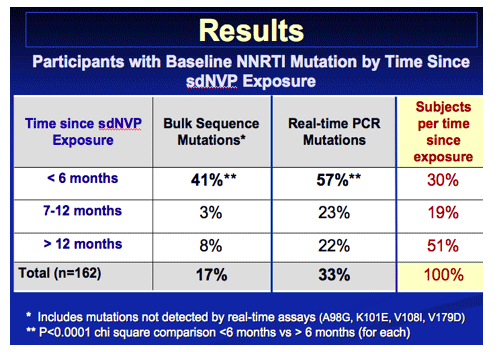
RESULTS: Bulk sequencing detected 15 mutations in 13 (5%) sdNVP-unexposed women, including four A98G, four V179D, two K101E, two K103N, and one each of V106M, V108I and G190A. Twenty-eight (17.5%) sdNVP-exposed women had sequence-detectable mutations: 20 at <6 months, one between 7 and 12 months, and seven >12 months after exposure (χ2 trend
P<0.0001). Sensitive testing detected variants of the four mutations assayed in 53/162 (33%) sdNVP-exposed women. SdNVP-exposed women with no detectable or only minority baseline mutations responded similarly to therapy (27% versus 18% failure; P=0.3). Among women with a sequence-detectable mutation at baseline, only 1/13 (0.8%) sdNVP-unexposed women experienced
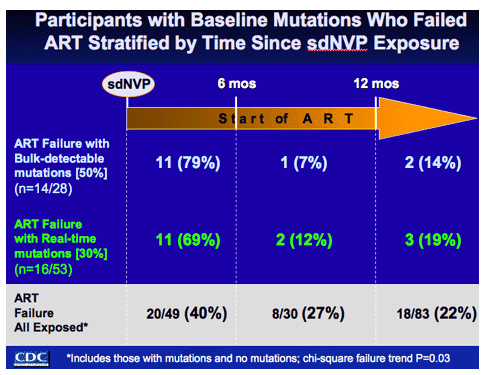
failure compared with 13/24 (54%) exposed women (P=0.01). For this latter group, the association with failure was strengthened by the subsets who also had minority mutations (8/13 failed) and those with A98G (3/4 failed). The four sdNVP-unexposed women with A98G did not experience failure. SdNVP-exposed women without detectable mutations were 1.8x more likely to fail than sdNVP-unexposed women (27% versus 15%; P=0.01). Regardless of detectable resistance, virological failure was greater (40%) when treatment was initiated <6 months after sdNVP (27% 7-12 months and 22% >12 months; χ2 trend P=0.03).
CONCLUSIONS: Although minority mutations when present with sequence-detectable mutations strengthened the association with failure, they alone did not predict failure. Even in the absence of detectable resistance, sdNVP-exposed women, when compared with unexposed, experienced significant virological failure close to the time of sdNVP exposure possibly suggesting transient resistance sanctuaries not readily detectable in peripheral viruses.





|
| |
|
 |
 |
|
|