 |
 |
 |
| |
Quest Tropism Assay: Performance of an HIV-1 Co-receptor Tropism Assay Utilizing Replicate V3 Loop Sequencing and Heteroduplex Analysis with Capillary Electrophoresis
|
| |
| |
R Baumann1, H Hamdan1, D Schwab1, T Robins2, J Vichayanonda1, and R Kagan1
1Quest Diagnostics Nichols Institute, San Juan Capistrano, CA; 2Quest Diagnostics Clinical Trials, Malibu, CA
Abstract
Introduction: CCR5 (R5) antagonist therapy may be ineffective in patients with preexisting CXCR4-tropic (X4) or dual/mixed virus. Genotypic tropism tests afford a more rapid and less complex means of screening for X4-tropic virus than phenotypic assays. In HIV samples from treatment-naïve patients, the SensiTrop II genotypic assay demonstrated 79% concordance with the standard Trofile phenotypic assay. We developed a genotypic tropism assay that couples replicate V3 loop sequencing with heteroduplex analysis by capillary electrophoresis (CE-HDA). Here we assess concordance of the CE-HDA method with the Trofile and SensiTrop II assays in samples from treatment-experienced patients.
Methods: Coreceptor tropism was assayed for 40 de-identified HIV-1 subtype B patient samples with genotypically predicted resistance to ≥4 reverse transcriptase or protease inhibitors by the Trofile (Monogram Biosciences, South San Francisco, CA), SensiTrop II (Pathway Diagnostics, Malibu, CA), and CE-HDA assays. A 141-bp HIV-1 envelope V3 loop amplicon was sequenced in duplicate, and tropism was predicted by bioinformatic analysis of amino acid sequences. For heteroduplex analysis, 4 FAM-labeled V3 loop R5 probes were annealed to the V3 amplicons and heteroduplexes were resolved on an ABI 3130xl CE analyzer.
Results: X4-utilizing virus was identified in 48% of samples by Trofile, 50% by SensiTrop II, and 56% by CE-HDA, but these differences were not statistically significant (McNemar test). CE-HDA was 72% concordant with Trofile (kappa=0.44). Sensitivity and specificity for X4 were 79% and 65% respectively. CE-HDA was 74% concordant with SensiTrop II (kappa=0.49) and SensiTrop II was 73% concordant with Trofile (kappa=0.45). Seven Trofile R5's were typed as X4 by CE-HDA, 4 of which were also typed as X4 by SensiTrop II. Four Trofile X4's were typed as R5 by CE-HDA, 2 of which were also R5 by SensiTrop II.
Summary: In samples from treatment-experienced patients, concordance between 2 independently developed genotypic tropism assays was comparable to the concordance of each assay with a phenotypic method. The proportion of X4 viruses detected did not vary significantly by assay type. Further comparisons of CE-HDA to the Trofile enhanced sensitivity (ES) phenotypic assay are warranted. Clinical outcome studies are needed to assess the predictive values of different tropism assays for patients treated with CCR5 antagonists.
Introduction
In patients with preexisting X4 virus, CCR5 antagonists may lead to the emergence of X4 variants and therapeutic failure.1,2 An assay for viral tropism is required prior to the initiation of treatment with CCR5 antagonists. A cell-based phenotypic assay (Trofile) was used in CCR5 antagonist clinical trials and is available commercially. Genotypic assays may afford a more rapid and less complex means of assessing tropism. A number of available genotypic methods to determine viral tropism target the V3 loop of the envelope gene, the primary HIV-1 tropism determinant.3
We have developed a genotypic assay that utilizes automated capillary electrophoresis heteroduplex analysis (CE-HDA) coupled with V3 loop sequencing and bioinformatic analysis. Here we assess the concordance of this method with both the Trofile phenotypic assay and the SensiTrop II genotypic assay in samples from treatment-experienced patients.
Method
Method Comparison Study
40 de-identified HIV-1 clinical samples (39 subtype B) with resistance to ≥4 RTI/PI were selected for analysis. The standard Trofile assay was performed at Monogram Biosciences. The SensiTrop II assay was performed at Pathway Diagnostics. Viral RNA was extracted from 0.5 mL plasma, and a 141-nt segment of the envelope gene spanning the V3 loop was reverse- transcribed and amplified by nested PCR. An additional 21 samples were also analyzed by the second-generation Trofile ES assay.
Heteroduplex Analysis
HDA mobility measures genetic distance, whereby targets more dissimilar to the R5 probes exhibit reduced mobility. An HDA assay based on previous methods was developed using FAM-labeled V3 probes derived from R5-tropic strains
JRCSF, SF162, SW54, and SW87.4-6
Heteroduplexes were resolved by capillary electrophoresis (CE) using ROX500 internal standards based on published methods.7 R5 and X4 mobility cutoffs were established by analysis of independently characterized specimens, and clinical samples were typed by sequencing.
DNA Sequence Analysis
V3 loop sequences were obtained by automated fluorescent sequencing. Nucleotide mixtures were translated into all possible amino acid permutations. Tropism was predicted using the 11/24/25 rule8-11 and the PSSMSI/NSI Position Specific Scoring Matrix.12
Results
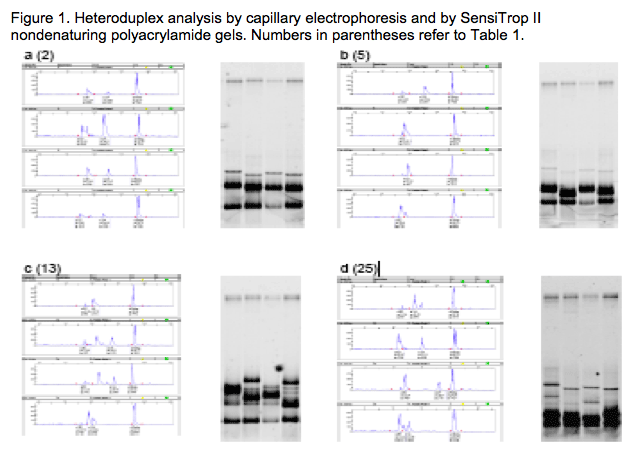
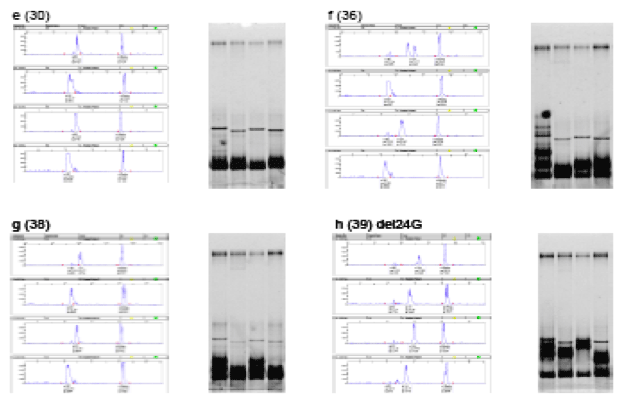
Both automated CE and nondenaturing polyacrylamide gel electrophoresis (PAGE) resolved V3 loop heteroduplexes (Fig 1). Use of ROX internal molecular weight standards in CE allowed for more precise determination of mobility cutoffs for R5:R5 and R5:X4 duplexes (Fig 1 a-h, red tickmarks). X4 heteroduplexes were correctly assigned by CE-HDA even when their mobility was close to the R5 cutoff (Fig. 1g and Table 1, sample 38). In some cases, X4 mobility shifts were only observed for a single probe (Fig. 1b and Table 1, sample 5).
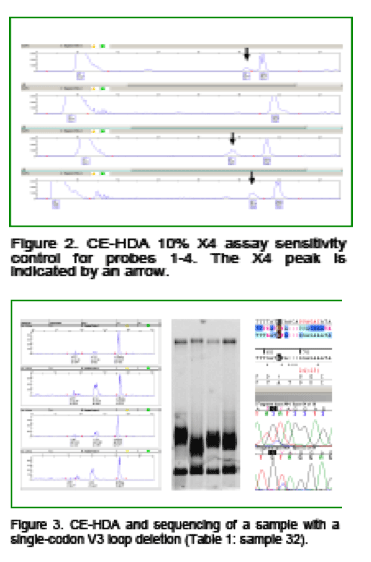
Population sequencing can detect minor variants present in 20%-30% of the viral population; HDA has greater sensitivity for minor variants.6 The ability of CE-HDA to detect minor X4 variants in R5 mixtures was assessed at total viral loads of 100,000 and 10,000 cp/mL at 10%, 5%, 2%, and 1% X4. X4 heteroduplexes at the 5% level were detected in 100% and 95% of 20 replicates at 100,000 cp/mL and 10,000 cp/mL, respectively. Consequently, a 90/10 R5/X4 mixed positive control was included in all assay runs to verify assay sensitivity (Fig 2).
Single-codon deletions (del24 or del23) were seen in 6/57 samples resulting in a mobility shift (Fig. 2h and Fig. 3). However, sequence analysis identified 5/6 such samples as R5-tropic in spite of the CE-HDA mobility shift; 4 of these were confirmed to be R5 by the Trofile assay (Table 1: samples 32, 39, 54 and 56; sample 15 was D/M by Trofile and sample 45 was X4 by both assays). Replicate sequences were concordant in all but 1 case (sample 31), where an X4 variant was seen in only 1 replicate. In 2 other cases, sequence data were obtained for only 1 of the 2 replicates.
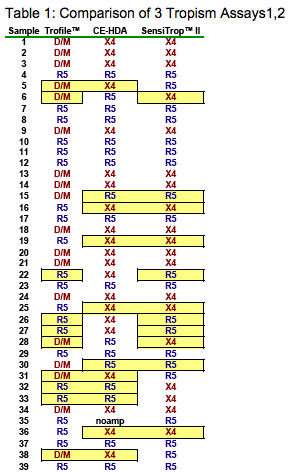
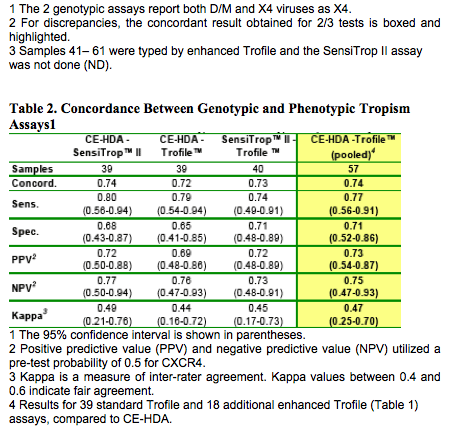
Comparison of 2 genotypic tropism assays to the standard Trofile assay showed agreement between all 3 assays for 23/39 samples; similar proportions of X4 virus were detected by SensiTrop II (50%), CE-HDA (56%) and Trofile (48%) (Table 1). Overall concordance in 2-way comparisons was 72%-74%, sensitivity for X4 was 0.74-0.80, and specificity was 0.65-0.71 (Table 2). Seven Trofile R5s were typed as X4 by CE-HDA, 4 of which were also typed as X4 by SensiTrop II (Table 1: 16, 19, 25 and 36). Four Trofile X4's were typed as R5 by CE-HDA, and 2 of these were also R5's in the SensiTrop II assay (Table 1: 15 and 30).
An additional 18 samples typed by the enhanced Trofile assay and by CE-HDA were in agreement in 14/18 cases, and results were pooled with the previous dataset (Table 1, samples 41-61). Overall concordance, sensitivity, and specificity for CE-HDA were 0.74, 0.77, and 0.71, respectively (Kappa statistic = 0.47, indicating a fair level of agreement) (Table 2).
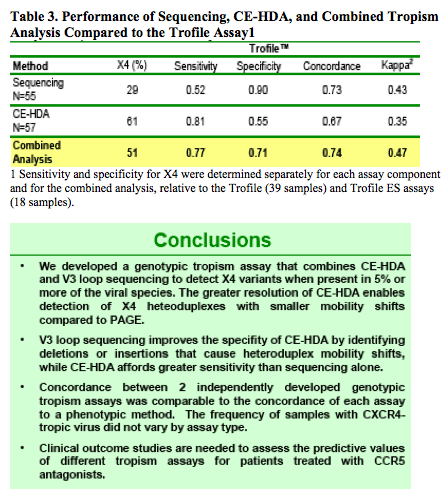
Recent reports showed improved sensitivity of population sequencing to detect X4 virus.10,11 To assess whether CE-HDA offers an advantage over population sequencing alone, we evaluated separately the concordances of the sequencing and CE-HDA to the Trofile results. Sequencing alone predicted X4 in only 29% of samples, compared to 61% for CE-HDA alone (McNemar Test: p<0.004; Table 3). Sequencing sensitivity for X4 was lower than that of the combined interpretation method (0.52 vs 0.77); CE-HDA alone had a slightly increased sensitivity (0.81), at the expense of reduced specificity (0.55 vs 0.71; Table 3).

REFERENCES
1. Lewis et al 2007. Antiviral Ther 12 (suppl 1):S65;
2. van der Ryst and Westby 2007. 47th ICAAC, abstract H-715;
3. Low et al 2008. AIDS Rev 10:143-51;
4. Delwart and Gordon. 1997. Methods 12:348-54;
5. Nelson et al 1997. J Virol 71:8750-8;
6. Weiser et al 2008. AIDS 22:469-79;
7. Davies et al 2006. Genomics 87:427- 32;
8. Cardozo et al 2007. AIDS Res Hum Retroviruses 23:415-26;
9. De Jong et al 1992. J Virol 66:6777- 80;
10. De Mendoza et al 2008. J Acquir Immune Defic Syndr 48:241-4;
11. Delobel et al 2007. J Clin Microbiol 45:1572-80;
12. Jensen et al 2003. J Virol 77:13376-88.
|
| |
|
 |
 |
|
|