 |
 |
 |
| |
No resistance to IDX184 was detected in 3-day and 14-day clinical studies of IDX184 in genotype 1-infected HCV subjects
|
| |
| |
Reported by Jules Levin EASL 2011 Berlin Germany March 30-Apr 2
Joseph F. McCarville, Gloria Dubuc, Eileen Donovan, Douglas Mayers, Maria Seifer, David N. Standring
Idenix Pharmaceuticals, Inc., Cambridge, MA, USA
AUTHOR CONCLUSIONS
In 3- and 14-day clinical trials with IDX184, no pre-existing S282T variant was detected.
The S282T resistance mutation was not detected following IDX184 treatment in either trial.
In the two clinical trials as well as in the preclinical studies, no other NS5B or other HCV mutation (N>1) has yet been identified as showing an association with IDX184 treatment.
In preclinical resistance selection studies in cultured cells, the S282T mutation took an average of over 7 weeks to appear and S282T replicon-bearing cells exhibited 5% of wild-type replication capacity.
Thus, the combined preclinical and clinical results are consistent with the concept of a high genetic barrier to resistance for nucleoside agents such as IDX184.
BACKGROUND
IDX184 is a liver-targeted 2'-C-methylguanosine nucleotide prodrug which inhibits the HCV NS5B polymerase of all genotypes in vitro. This work summarizes the resistance profile of IDX184 derived from early clinical studies
METHODS
Clinical Trials
Two double-blind studies were conducted in genotype 1 HCV-infected, treatment-naïve subjects
IDX-08C-003 was a 3-day proof-of-concept study with IDX184 administered as monotherapy to HCV-infected subjects.
33 subjects received once-daily doses of 25 mg, 50 mg, 75 mg or 100 mg IDX184; 8 subjects received matching placebo.
Samples were taken at baseline, at end of treatment (EOT) and 14 days post-treatment for NS5B population sequencing.
Two subjects with relatively poor virological response were selected for further examination by NS5B clonal sequencing-these subjects showed a modest on-treatment increase in viral load from nadir (0.5 to 0.7 log10 IU/mL) and had the weakest EOT responses to IDX184 in their respective cohorts (-0.23 and +0.1 log10 IU/mL compared to baseline).
Approximately 50 clones were obtained from samples from both subjects at baseline and after IDX184 dosing to determine if IDX184-resistant variants were present at levels too low to be detected by population sequencing.
IDX-08C-004 was a 14-day phase II study of IDX184 administered in combination with pegylated interferon and ribavirin to HCV-infected subjects.
65 subjects were assigned to receive 50 mg QD, 50 mg BID, 100 mg QD, 150 mg QD, 100 mg BID or 200 mg QD IDX184; 16 subjects received matching placebo. Two subjects dropped out of the trial after providing baseline samples.
Samples were taken at baseline, at EOT and 14 days post-treatment for NS5B population sequencing. None of the subjects experienced virologic breakthrough during treatment with IDX184, defined as a confirmed 1 log10 increase in HCV RNA from nadir or detectable HCV RNA in a subject with previously undetectable levels.
Clinical Analysis
RNA was isolated from all clinical samples with viral load ≥ 1,000 IU/mL using the Total Nucleic Acid isolation kit (Roche Applied Science). The NS5B domain was converted into cDNA by the Roche Transcriptor kit and amplified by nested PCR using the Expand High Fidelity kit.
For clonal sequencing, cDNA was ligated into a TOPO-TA based vector (Invitrogen) which was then used to transform bacterial cells. Positive clones were selected to inoculate LB cultures, and DNA plasmids were extracted for sequencing.
PCR products for population sequencing and plasmids for clonal sequencing were sequenced from both directions covering the entire NS5B region. For population sequencing, a mixed base was called if the lowest signal exceeded 25% of the highest signal.
Subject sequences were compared to consensus genotype 1a and 1b NS5B sequences derived from the Los Alamos National Laboratory (LANL) data base to assess the degree of baseline variability in subject sequences and to identify polymorphic sites within NS5B. NS5B positions were identified as polymorphic if they had ≥ 5% amino acid variability at that locus among the LANL sequences used to derive the consensus sequences.
In Vitro Analysis
Four cell lines containing genotype 1b HCV replicons were treated with IDX184 at concentrations ranging from 9X EC50 up to 50X EC50. Replicons were periodically sequenced to determine mutations that arose during exposure to IDX184.
The S282T mutation was introduced into the genotype 1b replicon using standard site-directed mutagenesis techniques.
In vitro resistance was determined by treating replicon-bearing cells with increasing concentrations of IDX184. EC50 values were calculated using XLfit© to fit dose-response curves.
RESULTS
IDX-08C-003 Clinical Resistance Results-NS5B Population Sequencing
In the 3-day study a total of 121 population NS5B sequences from the 33 IDX184-treated subjects and 8 placebo subjects in the IDX-08C-003 trial were examined (Table 1).
Baseline sequences showed significant variability relative to consensus genotype 1a and 1b NS5B sequences derived from the LANL data base.
At 51 residues where subject baseline sequences differed from a consensus NS5B sequence in more than one subject, 47 were identified as polymorphic sites
Analysis of EOT NS5B population sequences identified 53 changes in amino acid composition at 40 NS5B positions in subjects who received IDX184 (1.6 mutations/subject); 6 amino acid changes were observed in placebo subjects (0.8 mutations/subject). In all cases the observed changes involved mixed populations of amino acids.
Of the changes at 40 NS5B residues in subjects who received IDX184, 32 were observed in only one subject each, suggesting that the observed changes were random.
The other 8 residues at which a change was observed in more than one subject all involved mixtures of amino acids, and 5/8 were identified as being polymorphic sites
Subjects with changes at the 8 residues exhibited viral load reductions comparable to those without these mutations (-0.64 log10 IU/mL vs. -0.66 log10 IU/mL)
The difference between treatment and placebo groups in the frequency of amino acid changes from baseline to end-of-treatment was not statistically significant (p = 0.11).
No S282T mutation was detected by population sequencing in any baseline samples in IDX-08C-003. No S282T emerged following treatment with IDX184 or placebo.

IDX-08C-003 Clinical Resistance Results-NS5B Clonal Sequencing
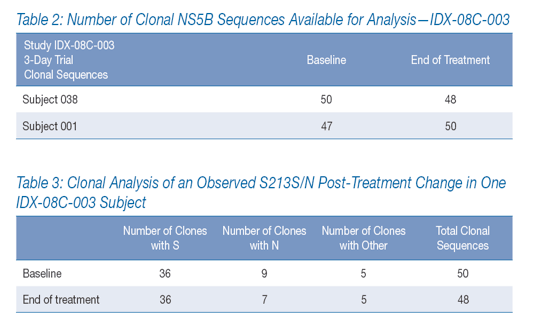
Approximately 200 clonal NS5B sequences were examined from the two IDX-08C-003 subjects with poor viral load response during IDX184 treatment-approximately 50 sequences from each subject at baseline and at EOT (Table 2).
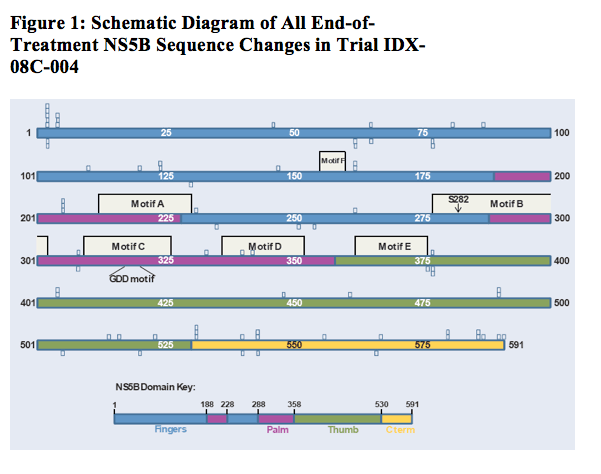
NS5B amino acids that changed at end of treatment relative to baseline are denoted by open boxes. The number of boxes at a given position indicates the number of changes observed. Boxes above the schematic protein represent changes in subjects receiving IDX184; boxes below the protein represent changes in placebo subjects. With only three exceptions, all changes involve mixtures of amino acids. Colored regions of the NS5B enzyme showing the fingers, palm, thumb and C-terminal domains follow O'Farrell et al. 20032. Highlighted motifs A through F identify regions conserved among RNA-dependent RNA polymerases3,4,5. Motif B is involved in nucleotide binding and includes S282, the site of the known IDX184 resistance mutation. The location of the GDD RNA polymerase active site motif is noted6.
·· A single amino acid change was observed by population sequencing in one of the two subjects: a change from S213 at baseline to mixed S/N213 at end of treatment. However, examination of the clonal sequences showed that the relative proportions of S213 and N213 remained essentially unchanged before and after treatment. Thus, the change reported by population sequencing simply reflected minor fluctuations in a mixed population (Table 3).
· Analysis of all clonal sequences from the two selected subjects did not detect the S282T mutation before or after IDX184 treatment. No other codon could be correlated with the relatively poor virological response of the subjects.


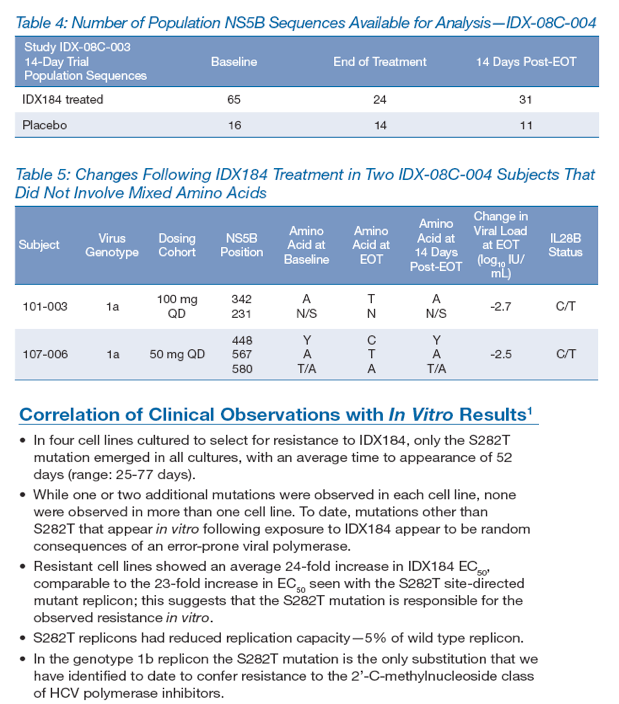
References
1. McCarville JF, Li B, Panzo RJ, et al. (2010) 5th International Workshop on Hepatitis C-Resistance and New Compounds. June 24-25, 2010; Boston.
2. O'Farrell D, Trowbridge R, Rowlands D, Jager J (2003) J. Mol. Biol. 326:1025-1035.
3. Koonin E (1991) J. Gen. Virol. 72:2197-2206.
4. Bressanelli S, Tomei L, Roussel A, et al. (1999) PNAS 96:13034-13039.
5. Lesburg C, Cable M, Ferrari E, et al. (1999) Nat. Struct. Biol. 6:937-943.
6. Munakata T, Nakamura M, Liang Y, et al. (2005) PNAS 102:18159-18164.
|
| |
|
 |
 |
|
|