 |
 |
 |
| |
Closing Summary by Jean-Michel Pawlotsky Part 1 Genetics, HBV Clinical Guidelines, HBV, HCV Fibrosis Progression, HBC Nucleotides, Alisporivir for HBV
|
| |
| |
EASL 2012 Sunday Morning Apr 22 Barcelona Spain
From Jules: I added the Program Abstract just below slides which include author & contact info.

PROGRAM ABSTRACT
Genome-wide association study identifies variants associated with liver fibrosis progression in HCV-infected patients
Speaker: Pierre-Yves Bochud Author: E. Patin1,2, Z. Kutalik3,4, J. Guergnon5, S. Bibert3, B. Nalpas2,6, E. Jouanguy1,2,7, M. Munteanu8, L. Bousquet2,9, L. Argiro10, P. Halfon11, A. Boland12, B. Mullhaupt13, D. Semela14, J.-F. Dufour15, M.H. Heim16, D. Moradpour3, A. Cerny17, R. Malinverni18, H. Hirsch16, G. Martinetti19, V. Suppiah20, G. Stewart20, D.R. Booth20, J. George20, J.-L. Casanova1,2,7, C. Bréchot21, C.M. Rice7, A.H. Talal22, I.M. Jacobson22, M. Bourliere23, I. Theodorou24, T. Poynard25, F. Negro26, S. Pol2,6, L. Abel1,2,7, P.-Y. Bochud3*, Swiss Hepatitis C Cohort Study, International Hepatitis C Genetics Consortium, French ANRS HC EP 26 Genoscan Study Group; EP and ZK, LA and PYB Contributed Equally to This Work Affiliation:
1INSERM U980, 2University Paris Descartes, Paris, France, 3University Hospital and University of Lausanne, 4Swiss Institute for Bioinformatics, Lausanne, Switzerland, 5Groupe Hospitalier Pitié Salpêtriere, 6Inserm U1016, Paris, France, 7Rockefeller University, New York, NY, USA, 8Biopredictive, 9INSERM U1016, Paris, 10INSERM-UMR 906, 11Hopital Ambroise Paré, Marseille, 12Centre National de Genotypage, Evry, France, 13University Hospital Zurich, Zurich, 14Kantonsspital St. Gallen, St. Gallen, 15University Hospital Bern, Bern, 16University Hospital Basel, Basel, 17Clinica Moncucco, Lugano, 18Pourtales Hospital, Neuchatel, 19Institute for Medical Microbiology, Bellinzona, Switzerland, 20University of Sydney, Sydney, ACT, Australia, 21INSEM U785, Villejuif, France, 22Weill Cornell Medical College, New York, NY, USA, 23Hôpital St Joseph, Marseille, 24INSERM UMR-S 945, 25Hôpital Pitié Salpêtriere, Paris, France, 26University Hospital Geneva, Geneva, Switzerland. *pierre-yves.bochud@chuv.ch
Background: Clearance of hepatitis C virus (HCV) infection is modulated by IL28B polymorphisms, as shown by genome-wide association studies (GWAS). Only a fraction of patients with chronic infection develop liver fibrosis, a process that may also be controlled by human genetic factors. We carried out a two-stage GWA study of liver fibrosis progression related to HCV infection.
Patients and methods: We studied well characterized HCV-infected patients of European descent who had liver biopsy before treatment. We defined various liver fibrosis phenotypes on the basis of Metavir score, with and without taking the duration of HCV infection into account. GWAS was conducted on a filtered primary cohort of 1,161 patients, using 780,650 single nucleotide polymorphisms (SNPs). We genotyped 96 SNPs with P-values< 5x10-5 in an independent replication cohort of 962 patients. Finally, we assessed the most interesting replicated SNPs in an additional sample of 219 patients genotyped in a previous GWAS on HCV clearance.
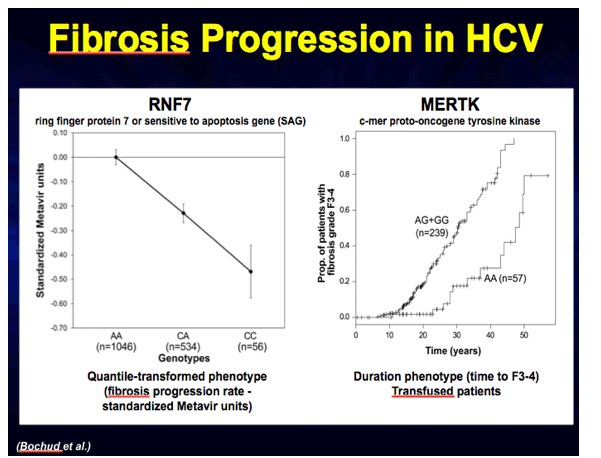
Results: In the combined cohort of 2,342 HCV-infected patients, two SNPs, one in the total sample and one in blood-transfused patients, provided genome-wide significant evidence of association with fibrosis progression (P combined=8·9x10-9 and 2·1x10-9, respectively). The first SNP is located within RNF7, which acts as an antioxidant protecting against apoptosis. The other, together with another replicated SNP, (P combined=5·4x10-7), are linked to two functionally related genes, MERTK and TULP1, involved in phagocytosis of apoptotic cells by macrophages.
Conclusion: Our GWAS identified several susceptibility loci for HCV-induced liver fibrosis related to the apoptosis pathway, providing new insights into the mechanisms underlying fibrosis development and paving the way for novel therapeutic strategies.
PROGRAM ABSTRACT
Pegylated-interferon-alpha alone or in combination with entecavir restores ISG responsiveness and reduces intrahepatic viral loads and antigenemia in hepatitis B virus infected humanized mice Speaker: Lena Allweiss Author: L. Allweiss1*, M. Lütgehetmann1,2, T. Volz1, T. Bornscheuer1, A.W. Lohse1, J. Petersen3, H. Ma4, K. Klumpp4, S. Fletcher4, M. Dandri1 Affiliation: 1Internal Medicine, 2Medical Microbiology, Virology & Hygiene, University Medical Center Hamburg-Eppendorf, 3Liver Centre, Asklepiosklinik St. Georg, Hamburg, Germany, 4Hoffmann-La Roche, Nutley, NJ, USA. *lenaallweiss@hotmail.com
Background: The direct antiviral efficacy of interferon alpha (IFN-α) treatment on hepatic hepatitis B virus (HBV) replication has not been well defined. Using HBV-infected humanized uPA/SCID mice we recently showed that treatment with conventional IFN-α induced short-lived (< 24 hours) suppression of HBV replication and the responsiveness of infected human hepatocytes to IFN-α was impaired (Gastroenterology 2011:140;2074-83). Aim of this study was to investigate whether the use of pegylated interferon-α (peg-IFN-α), alone or in combination with entecavir (ETV), could induce more effective antiviral control in HBV-infected mice and improve hepatocyte responsiveness to IFN-α.
Methods: HBV-infected humanized uPA/SCID mice displaying median viral titers of 3.6x10E8 HBV-DNA/ml (n=23) received either peg-IFN-α monotherapy (25ng/gr, twice/week; n=6), ETV monotherapy (100ng/gr/day; n=6), or both drugs in combination (n=6), while 5 animals were left untreated as controls. Serological changes were determined prior to, and after 2 and 4 weeks of treatment. Intrahepatic analyses were performed by qRT-PCR and immunohistochemistry.
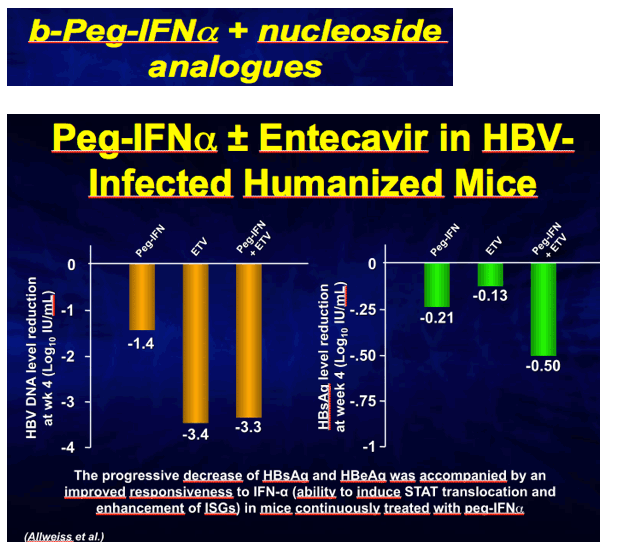
Results: After 4 weeks of therapy, the median reduction in viremia was 1.4Log, 3.4Log and 3.3Log in mice treated with peg-IFN-α, ETV or the combination thereof, respectively. Despite the larger viremia reduction provided by ETV, the reduction of intrahepatic HBcAg expression levels was more pronounced in mice receiving peg-IFN-α alone or combination therapy, than in mice receiving ETV monotherapy. Peg-IFN-α alone or in combination also reduced the levels of circulating HBsAg (0.21Log; 0.5Log) and HBeAg (0.55Log; 0.66Log), whereas the median decrease in these viral antigens was less pronounced in mice receiving ETV monotherapy (HBsAg=∼0.13Log; HBeAg=∼0.24Log). Remarkably, the progressive decrease of viral antigens was accompanied by an improved responsiveness to IFN-α, which was demonstrated by the ability to induce STAT translocation and enhancement of human interferon stimulated genes (ISG) in mice continuously treated with peg-IFN-α.
Conclusions: The strong reduction of intrahepatic viral loads and antigenemia achieved in vivo in the absence of adaptive immune responses emphasizes the qualitatively different antiviral effects induced by peg-IFN-α compared to polymerase inhibitors, thus providing a rationale for exploring the potential of therapeutic strategies based on the combination of distinct types of antiviral agents.
PROGRAM ABSTACT
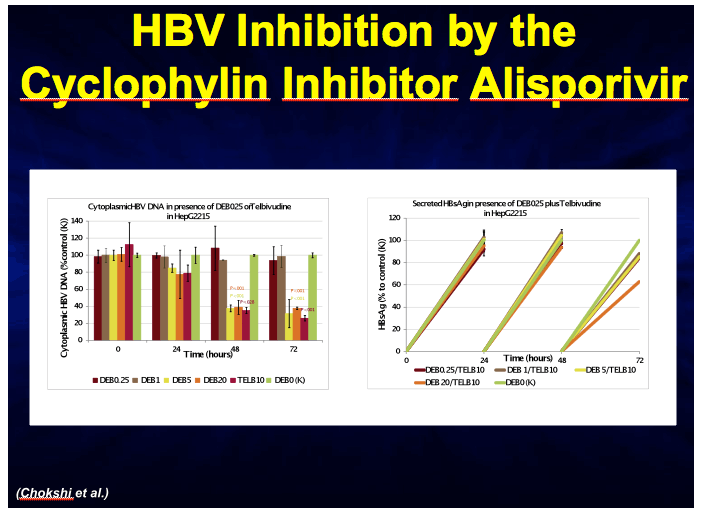
ALISPORIVIR-INDUCED INHIBITION OF CELLULAR CYCLOPHILINS DISRUPTS HEPATITIS B VIRUS (HBV) REPLICATION IN VITRO AND IS SYNERGISTIC IN COMBINATION WITH DIRECT ANTIVIRAL TARGETING HBV-DNA POLYMERASE
Speaker: Shilpa Chokshi Author: S. Phillips1, S. Chokshi1*, A. Riva1, N.V. Naoumov2 Affiliation: 1Foundation for Liver Research, Institute of Hepatology, London, UK, 2Novartis Pharma AG, Basel, Switzerland. *s.chokshi@researchinliver.org.uk
Background/aims: Cyclophilins are intracellular proteins with enzymatic activity - peptidyl-prolyl-isomerase that plays a major role in the life cycle of Hepatitis C virus. By targeting host cyclophilins Alisporivir (DEB025) exerts potent anti-HCV activity in vitro and in clinical studies. We have recently shown in vitro that cyclophilin inhibition with Alisporivir or NIM811 also interferes with HBV replication, with Alisporivir having a greater effect than NIM811. To elucidate the underlying mechanisms, in the present study we compared in vitro the effects on HBV replication of Alisporivir alone, Alisporivir in combination with a potent antiviral targeting HBV-DNA polymerase, and in cells after selective knockdown of individual cyclophilins.
Material and methods: Stably(HepG2215) and transiently(HUH-7) transfected cells, producing full HBV virions and HBsAg particles, were treated with a range of Alisporivir concentrations (0.25/1.0/5.0/20 ug/ml) alone, Telbivudine alone, or combinations of Alisporivir and Telbivudine. To determine the involvement of individual cyclophilins, HepG2215 cells were transfected with siRNA-specific for cyclophilin (Cyp) A, C or D and additionally treated with Alisporivir. Cytoplasmic extracts and supernatants were harvested at baseline; 24, 48 and 72 hours post-treatment. The kinetics of antiviral activity was assessed by quantitation of intracellular and secreted HBV-DNA (real-time qPCR) and HBsAg levels (ELISA).
Results: Both in HepG2215 and HUH-7 cells, Alisporivir treatment resulted in dose-dependent reduction of intracellular and secreted HBV-DNA at all time points, by 70% (p=0.004) and 63% (p< 0.001), respectively, compared with untreated controls. The combination of Alisporivir and Telbivudine had greater effects in reducing intracellular (p=0.001) and secreted (p=0.028) HBV-DNA, and >3-fold reduction of HBsAg versus either Alisporivir or Telbivudine alone. CypA, C or D expression was markedly reduced after transfection with corresponding siRNA, which was associated with significant decrease of HBV-DNA and HBsAg levels (p< 0.001). Alisporivir treatment of cells silenced for CypA, C or D further reduced HBV-DNA and HBsAg levels, with greater antiviral effects in CypC or CypD silenced cells, compared with CypA silenced cells (p< 0.001).
Conclusion: These results suggest that Alisporivir interferes with multiple sites of HBV replication and its antiviral activity is synergistic with direct antiviral targeting viral DNA polymerase, such as Telbivudine.
|
| |
|
 |
 |
|
|