| |
Analysis of long-term persistence of resistance mutations within the hepatitis C virus NS3 protease after treatment with telaprevir or boceprevir
|
| |
| |
Download the PDF here
Journal of Clinical Virology 52 (2011) 321- 327
Simone Sussera , Johannes Vermehrena , Nicole Forestiera , Martin Walter Welkera , Natalia Grigorianb ,
Caterina Fuller a, Dany Pernera, Stefan Zeuzema, Christoph Sarrazina,
a Klinikum der Goethe Universitαt, Medizinische Klinik 1, Theodor-Stern-Kai 7, 60590 Frankfurt am Main, Germany
b Universitαtsklinikum Homburg, Klinik fur Innere Medizin 2, Kirrberger Str., Homburg/Saar, Germany
Abstract
Background
Telaprevir and boceprevir are highly selective hepatitis C virus (HCV) NS3/4A proteaseinhibitors in phase 3 development. Viral breakthrough during mono- and triple-therapies with PEG-interferon and ribavirin and relapse is associated with resistance.
Objectives
Potential persistence of resistance mutations during long-term follow-up should be analyzed.
Study design
Clonal sequence analysis of the NS3-protease gene was performed at long-term follow-up in HCV genotyp-1 infected patients who received telaprevir or boceprevir within phase-1b studies for comparison with resistant variants present directly after the end-of-treatment.
Results
After a median follow-up of 4.2 years in 28 of 82 patients HCV-RNA was still detectable. Resistance variants were detected in two of 14 telaprevir- and in four of 14 boceprevir-treated patients. For telaprevir patients two low-level (V36M, V36A) and one high-level (A156T) mutation associated with resistance were detected at low frequencies (4-9% of the clones). In five boceprevir-treated patients four low level mutations (V36A, T54A/S, V55A) were observed at low frequencies (1-10%) while in one patient additionally a combined variant (T54S + R155K) was detected at 94%. Presence of resistant variants at long-term follow-up was not predictable by variants detected at the end-of-treatment. In one patient a V55A variant which was dominant already at baseline was still detectable at long-term follow-up.
Conclusions
In the majority of patients after short-term treatment with telaprevir or boceprevir wild-type NS3-protease isolates are detectable by clonal sequencing at long-term follow-up. Detectable resistance mutations in single patients are not predictable by initial frequencies of variants.
Abbreviations
· HCV, hepatitis C virus;
· NS, non-structural protein;
· RNA, ribonucleic acid;
· WHO, World Health Organization;
· DAA, directly acting antiviral agent;
· TID, every 8 h;
· BID, every 12 h;
· wk, week;
· QID, every 6 h;
· PEG-IFNα, pegylated interferon-alpha;
· PCR, polymerase chain reaction;
· GT, genotype;
· bp, base pairs;
· SOC, standard-of-care;
· IU, international units;
· HIV, human immunodeficiency virus;
· HBV, hepatitis B virus;
· SVR, sustained virologic response
1. Background
Chronic hepatitis C virus (HCV) infection is a serious public health problem affecting an estimated 130 million people worldwide.1 The current standard-of-care, pegylated interferon-alpha (PEG-IFN) plus ribavirin, is of limited efficacy with eradication of the virus in approx. 50% of patients.2 A large number of directly acting antiviral agents (DAA) targeting the nonstructural-(NS)3-protease, the NS5A-protein, the RNA-dependent RNA-polymerase NS5B, as well as host cell proteins are currently in phase-1 to -3 clinical trials. [3] and [4] Telaprevir and boceprevir, the most advanced agents, have completed phase 3 studies. Results of clinical studies have shown a significant improvement of sustained virologic response (SVR) rates with triple therapies with telaprevir or boceprevir in combination with PEG-IFN and ribavirin in treatment-naïve and -experienced HCV genotype-1 infected patients. [5], [6], [7] and [8]
For monotherapy with NS3-protease, NS5A-protein and non-nucleoside polymerase inhibitors rapid selection of resistant variants has been observed, [9] and [10] which represents one of the major problems of DAAs. Combination therapy with PEG-IFN and ribavirin or with different DAAs has been shown to reduce the likelihood of virologic break-through due to resistance developing. [7], [8], [9], [10], [11], [12], [13] and [14] However, even using triple therapy with pegylated interferon, ribavirin and NS3 proteaseinhibitors, still a significant number of patients experience viral breakthrough in association with selection of drug-resistant variants. [7], [8], [11], [12] and [14] Furthermore, variants associated with resistance have been detected in patients with relapse after the end-of-treatment. [11], [12] and [14] Finally, viral variants known to confer resistance to NS3-protease and NS5B non-nucleoside inhibitors have been detected in 0.2-2.8% of patients as the pre-existing dominant strain [15], [16], [17] and [18] and preexisting R155K variants seem to be associated with a reduced response to boceprevir and telaprevir. [19] and [20]
2. Objectives
Preexisting or selected variants may affect virologic response to DAA. Here, we present long-term follow-up data on patients who were enrolled in phase-1 studies with telaprevir or boceprevir. After a median follow-up of 4.23 years clonal sequence analysis was performed in 28 patients with detectable HCV-RNA for analysis of potential persistence of viral variants (at amino acid (aa) positions 36, 54, 55, 155, 156, and 170) previously described to confer resistance to boceprevir or telaprevir. [21], [22], [23], [24], [25], [26] and [27]
3. Study design
3.1. Patient population
Altogether 82 patients with chronic HCV genotyp-1 infection were enrolled in phase-1 clinical trials with telaprevir and boceprevir at Saarland University Hospital. Up to 5.5 years after termination of study treatment 42/82 patients could be contacted for a long-term follow-up visit. HCV-RNA was still detectable in 34/42 patients (n = 6 placebo; n = 14 telaprevir; n = 14 boceprevir).
3.2. Telaprevir
Thirty-three HCV genotyp-1 infected patients were enrolled into two randomized, double-blind, placebo-controlled phase-1b trials, which were described recently. [28] and [29] The patients were treated with telaprevir alone (450 mg TID, 750 mg TID, or 1250 mg BID) or in combination with PEG-IFN-alpha-2a (180 μg/wk) for 2 weeks (750 mg TID). Clonal sequence analysis of the NS3-protease gene could be performed in 14 available patients with ongoing HCV-infection during long-term follow-up. Clonal resistance analysis at end-of-treatment was obtained from a previous study.21
3.3. Boceprevir
Forty-nine HCV genotyp-1 infected patients were enrolled into randomized, double-blind, placebo-controlled phase-1b trials. [30] and [31] The patients were treated with boceprevir alone (200 mg BID, 400 mg BID, or 400 mg TID) or in combination with PEG-IFN-alpha-2b (1.5 μg/kg body weight/wk) for 2 weeks (400 mg TID, 400 mg QID, or 600 mg QID) or they were randomized to different sequences of three periods of treatment with boceprevir-monotherapy (200 mg or 400 mg TID) for 7 days, PEG-IFNα-2b-monotherapy once weekly for 14 days and combination of the two for 14 days. Between the different treatment schedules therapy was interrupted for 14 days. Clonal sequence analysis of the NS3-protease gene could be performed in 14 available patients with ongoing HCV-infection at long-term follow-up. Clonal resistance analysis was performed in all patients at end-of-treatment in the present study or obtained from previous studies (patients #3, #9, #13). [27] and [32]
Methods used in this and the former studies were the same, regarding RNA isolation, RT-PCR, amplification, sequencing, and analyzing sequences.
Written informed consent was obtained from each patient in accordance with the 1975 Declaration of Helsinki.
3.4. HCV-RNA extraction
Viral RNA was extracted from 140 μL serum using the QIAamp Viral RNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions. RNA quality was assessed by calculating the absorbance ratio OD260nm/280nm using NanoDrop model ND-1000 (PeqLab, Erlangen).
3.5. Amplification and sequencing of the HCV NS3-protease gene
The complete region encoding the NS3-protease was amplified by semi-nested RT-PCR by applying 8 μL of the isolated RNA. The appropriately sized PCR-products were purified using the QIAquick Gel Extraction Kit (Qiagen, Hilden, Germany) and subsequently cloned using the StrataClone PCR Cloning Kit (Agilent Technologies, La Jolla, CA).
Isolated plasmid DNA from the molecular clones was subjected to sequence-PCR according to the manufacturer's instructions using the M13-forward or -reverse primers (BigDye Deoxy Terminators; Applied Biosystems, Foster City, CA). Sequencing was performed by the 3130xl Genetic Analyzer (Applied Biosystems, Foster City, CA).
Baseline consensus sequences were obtained in the context of the phase-1b studies and act as reference for analysis of the long-term follow-up samples. Baseline consensus sequences were checked for mutations associated with resistance at aa positions 36, 54, 55, 155, 156, and 170 as well.
3.6. Sequence analysis
Sequences from the N-terminal 181 aa of the HCV NS3-protease were aligned and analyzed for mutations which were known to confer resistance against telaprevir and/or boceprevir. The sequences were analyzed using the software Mutational Surveyor (SoftGenetics, State College, PA). Phylogenetic analyses were performed with the ClustalW-Multiple Sequence Alignment tool (www.ebi.ac.uk/Tools/msa/clustalw2). The cut-off values for sensitivity of the sequence analysis represent the percentage corresponding to 1 clone out of the number of clones sequenced.
4. Results
4.1. Clinical outcome of patients
4.1.1. Telaprevir
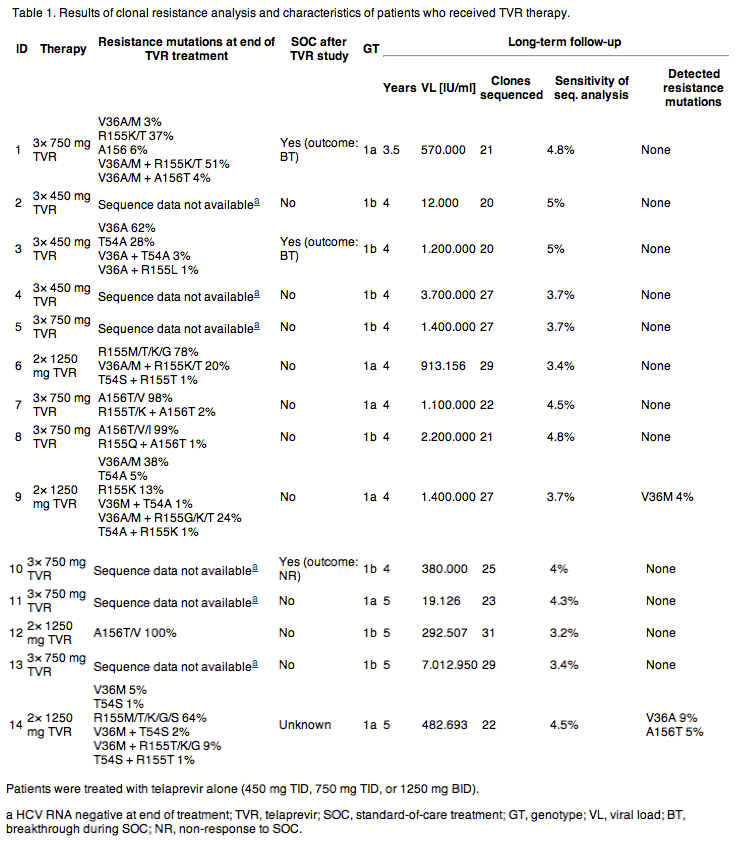
Twenty-one patients presented for a long-term follow-up visit 3.5-5 (mean 4.25 ± 0.5) years after termination of telaprevir therapy. Seven patients attained SVR after SOC in the meantime. Characteristics of the remaining patients are summarized in Table 1.
4.1.2. Boceprevir
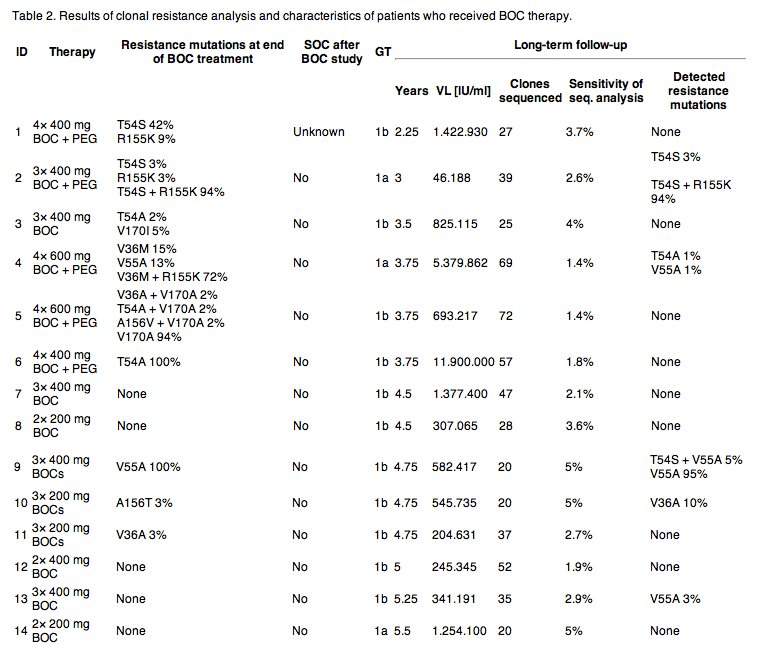
Fifteen patients were available for a long-term follow-up visit 2.25-5.5 (mean 4.2 ± 0.9) years after direct antiviral therapy with boceprevir. One of them achieved SVR with SOC after boceprevir dosing. Thus, the HCV NS3-protease gene of 14 patients could be analyzed (Table 2).
4.2. Resistance analysis
4.2.1. Telaprevir
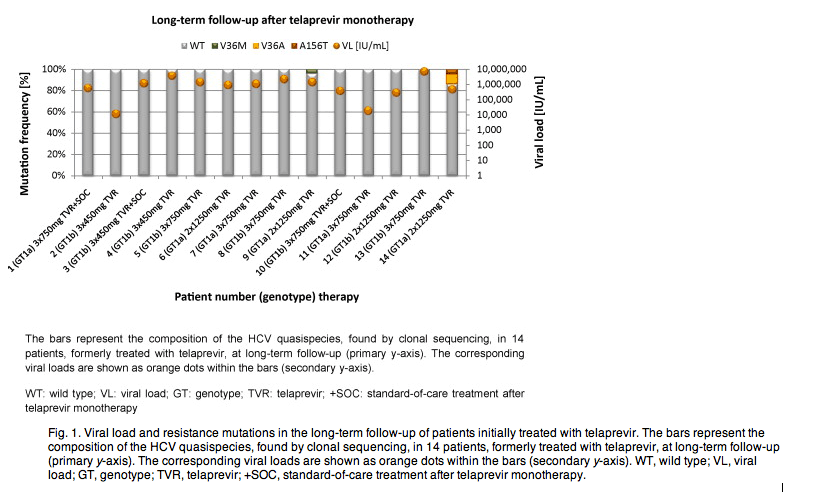
Clonal sequencing at long-term follow-up revealed only wild-type NS3-protease variants in 12 patients. One subtype 1a infected patient (#9, 2x 1250 mg) showed the V36M variant at 4% frequency, known to confer low level resistance to telaprevir. In another subtype 1a patient (#14, 2x 1250 mg) 9% of the analyzed clones showed V36A and even A156T, conferring the highest resistance against telaprevir, was present in 5% of the quasispecies. In patient #9 V36M was already detectable at the end of telaprevir-monotherapy, while patient #14 did not exhibit V36A nor A156T after telaprevir treatment. Both patients with NS3 proteaseinhibitor resistance variants were infected with HCV subtype 1a. Patient #9 did not receive SOC between completion of the telaprevir phase-1b study and long-term follow-up, for patient #14 this information is unknown (Table 1, Fig. 1).
4.2.2. Boceprevir
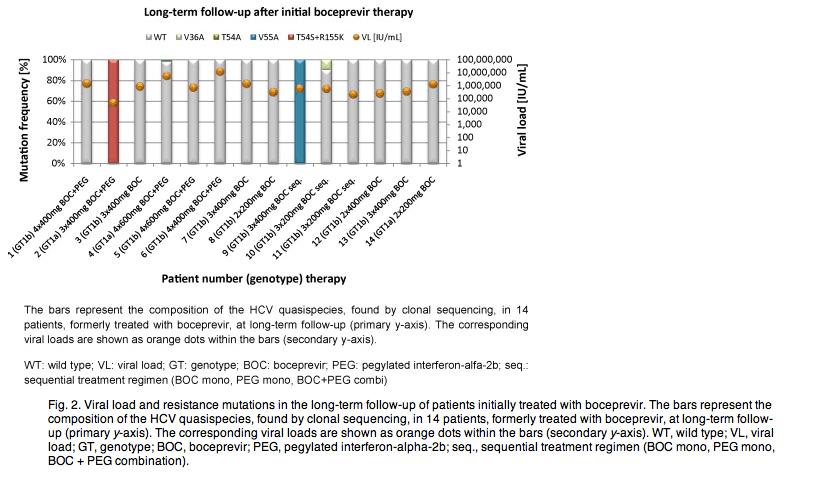
In patient #2 (GT1a; 3x 400 mg boceprevir + PEG) 94% of all clones contained the T54S + R155K double mutation and 3% contained T54S as single variant 3 years after proteaseinhibitor treatment. Directly after EOT T54S + R155K was present in 94% together with 3% T54S and 3% R155K as single mutations. In patient #4 (GT1a), 3.75 years after receiving 600 mg boceprevir QID plus PEG-IFN 1% T54A and 1% V55A were detected, each on separate clones (cut-off 1.4%). Contrary to V55A (13% frequency at EOT) T54A could not be detected after the end-of-treatment. V36M (EOT: 15%) and V36M + R155K (EOT: 72%) decreased completely to undetectable levels (<1.4%) during the 3.75 years. In patient #9 (GT1b), after 4.75 years we found 95% V55A and 5% T54S + V55A and in patient #10 (GT1b) 10% V36A, both were treated sequentially with boceprevir mono for 7 days (patient #9: 3x 400 mg; patient #10: 3x 200 mg), PEG mono and boceprevir + PEG in combination for 14 days, respectively. V55A which was detected in 100% of the clones from patient #9, was already present at baseline and at the end-of-treatment as natural and only variant with a frequency of 100%.32 Directly after boceprevir dosing, patient #10 exhibited only A156T (3%), which was no longer detectable (cut-off: 5%) in the long-term follow-up analysis. Patient #13 (GT1b) showed 3% V55A (cut-off: 2.9%) 5.25 years after boceprevir therapy (3x 400 mg), while no variants were detected after EOT.
The remaining nine patients showed no resistant variants above the limit of detection, despite detectable resistance mutations at the end of DAA treatment with boceprevir alone or in combination with PEG-interferon-alpha (patients #1, #3, #5, #6, and #11; Table 2, Fig. 2).
4.3. Phylogenetic analysis
With phylogenetic analyses we confirmed that the long-term follow-up sequences are closely related to the baseline and end-of-treatment sequences. The consensus sequences within one patient at these time points show an accordance of at least 95%, whereas the consensus sequences of different patients show identities down to 81% (Fig. 3 and Fig. 4).
See pdf
Fig. 3. Phylogenetic analysis of the consensus sequences from baseline, end-of-treatment, and long-term follow-up of each patient treated with telaprevir.
Fig. 4. Phylogenetic analysis of the consensus sequences from baseline, end-of-treatment, and long-term follow-up of each patient treated with boceprevir.
5. Discussion
HCV is a positive-stranded RNA virus with replication without a DNA intermediate and no mechanisms are known for HCV-RNA to be archived. [33] and [34] Due to the short half-life, the rapid turnover and the low fidelity of the HCV NS5B RNA-dependent RNA-polymerase a high number of viral variants is generated with production of all possible single and double variants every day. [35] and [36] Indeed, highly sensitive clonal sequence analyses showed a rapid selection of resistant variants during the first days of treatment with the HCV NS3 proteaseinhibitors telaprevir and boceprevir but also a rapid disappearance of resistant variants shortly after withdrawal of direct antiviral treatment. [21], [22] and [27] However, mutations within the NS3-protease conferring resistance to proteaseinhibitors do not impair infectivity of HCV virions and thus may persist at low levels also during long-term follow-up. In the present study, in the majority of patients only wild-type NS3-protease isolates were detectable 2.25-5.5 (mean 4.23 ± 0.7) years after the end of telaprevir or boceprevir treatment by clonal sequence analysis. Only in two patients with initial telaprevir treatment (1250 mg BID) and in five patients with initial boceprevir treatment (600 mg QID, 400 mg and 200 mg TID, respectively) resistant variants have been detected by clonal sequencing at long-term follow-up. V36M in patient #9 seems to be persisting since its selection during telaprevir monotherapy where it was detectable at end-of-treatment already. Initial sequencing of patient #14 was obtained at EOT after a viral breakthrough during telaprevir-monotherapy and V36A/A156T were not detectable at this time point.21 However, it could be possible, that V36A and A156T, which were detected by clonal analysis 5 years later, were present at earlier timepoints during telaprevir treatment. A156T has been observed as natural variant in the liver of an untreated patient and thus so far unknown compensatory mechanisms explaining the relatively high replication capacity of this variant within the HCV quasispecies may exist.37 Interestingly, for boceprevir and telaprevir resistance variants, the mutational pattern detected at long-term follow-up could not generally be predicted from variants present at the end-of-treatment which most likely is explained by the highly dynamic changes of HCV quasispecies.
Our study has several limitations: Boceprevir was dosed at lower levels than in phase-2/3 studies. Furthermore, during phase-1b studies all patients received only short-term treatment with boceprevir or telaprevir. Longer treatment durations and especially a continuous dosing of the proteaseinhibitor after viral breakthrough or persistent HCV replication during triple therapies with PEG-interferon and ribavirin might increase the probability for selection of resistance variants. Such variants together with compensatory mutations may have a higher replicative fitness and thus a higher likeliness for long-term persistence at significant frequencies within the HCV-quasispecies. Finally, clonal sequence analysis performed in the initial phase-1 studies as well as in the present study have limited sensitivities to detect resistance mutations present at low frequencies. Further studies on potential persistence of resistant variants by deep-sequencing methods are required to complete our understanding of viral turnover of these variants.
Taken together, we have shown a rapid decline of the number of patients with NS3-protease resistance mutations as well as the frequency of resistant viral variants within the HCV quasispecies from the initial months after the end of short treatment intervals until long-term follow-up of up to 5.5 years. Future studies based on clonal resistance analysis or even deep-sequencing should be performed after failure to full-course telaprevir and boceprevir treatment regimens.
|
|
| |
| |
|
|
|